Direct submissions to PCI Ecology from bioRxiv.org are possible using the B2J service
Latest recommendations

| Id | Title * | Authors * | Abstract * | Picture * | Thematic fields * | Recommender | Reviewers | Submission date▲ | |
|---|---|---|---|---|---|---|---|---|---|
05 Jun 2024
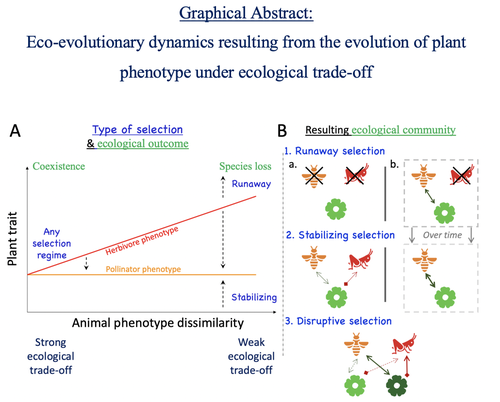
Attracting pollinators vs escaping herbivores: eco-evolutionary dynamics of plants confronted with an ecological trade-offYoussef Yacine, Nicolas Loeuille https://doi.org/10.1101/2021.12.02.470900Plant-herbivore-pollinator ménage-à-trois: tell me how well they match, and I'll tell you if it's made to lastRecommended by Sylvain Billiard based on reviews by Marcos Mendez and Yaroslav Ispolatov based on reviews by Marcos Mendez and Yaroslav Ispolatov
How would a plant trait evolve if it is involved in interacting with both a pollinator and an herbivore species? The answer by Yacine and Loeuille is straightforward: it is not trivial, but it can explain many situations found in natural populations. Yacine and Loeuille applied the well-known Adaptive Dynamics framework to a system with three interacting protagonists: a herbivore, a pollinator, and a plant. The evolution of a plant trait is followed under the assumption that it regulates the frequency of interaction with the two other species. As one can imagine, that is where problems begin: interacting more with pollinators seems good, but what if at the same time it implies interacting more with herbivores? And that's not a silly idea, as there are many cases where herbivores and pollinators share the same cues to detect plants, such as colors or chemical compounds. They found that depending on the trade-off between the two types of interactions and their density-dependent effects on plant fitness, the possible joint ecological and evolutionary outcomes are numerous. When herbivory prevails, evolution can make the ménage-à-trois ecologically unstable, as one or even two species can go extinct, leaving the plant alone. Evolution can also make the coexistence of the three species more stable when pollination services prevail, or lead to the appearance of a second plant species through branching diversification of the plant trait when herbivory and pollination are balanced. Yacine and Loeuille did not only limit themselves to saying "it is possible," but they also did much work evaluating when each evolutionary outcome would occur. They numerically explored in great detail the adaptive landscape of the plant trait for a large range of parameter values. They showed that the global picture is overall robust to parameter variations, strengthening the plausibility that the evolution of a trait involved in antagonistic interactions can explain many of the correlations between plant and animal traits or phylogenies found in nature. Are we really there yet? Of course not, as some assumptions of the model certainly limit its scope. Are there really cases where plants' traits evolve much faster than herbivores' and pollinators' traits? Certainly not, but the model is so general that it can apply to any analogous system where one species is caught between a mutualistic and a predator species, including potential species that evolve much faster than the two others. And even though this limitation might cast doubt on the generality of the model's predictions, studying a system where a species' trait and a preference trait coevolve is possible, as other models have already been studied (see Fritsch et al. 2021 for a review in the case of evolution in food webs). We can bet this is the next step taken by Yacine and Loeuille in a similar framework with the same fundamental model, promising fascinating results, especially regarding the evolution of complex communities when species can accumulate after evolutionary branchings. Relaxing another assumption seems more challenging as it would certainly need to change the model itself: interacting species generally do not play fixed roles, as being mutualistic or antagonistic might generally be density-dependent (Holland and DeAngelis 2010). How would the exchange of resources between three interacting species evolve? It is an open question. References Fritsch, C., Billiard, S., & Champagnat, N. (2021). Identifying conversion efficiency as a key mechanism underlying food webs adaptive evolution: a step forward, or backward? Oikos, 130(6), 904-930. Yacine, Y., & Loeuille, N. (2024) Attracting pollinators vs escaping herbivores: eco-evolutionary dynamics of plants confronted with an ecological trade-off. bioRxiv 2021.12.02.470900; doi: https://doi.org/10.1101/2021.12.02.470900 | Attracting pollinators vs escaping herbivores: eco-evolutionary dynamics of plants confronted with an ecological trade-off | Youssef Yacine, Nicolas Loeuille | <p style="text-align: justify;">Many plant traits are subject to an ecological trade-off between attracting pollinators and escaping herbivores. The interplay of both plant-animal interaction types determines their evolution. As most studies focus... |  | Eco-evolutionary dynamics, Herbivory, Pollination, Theoretical ecology | Sylvain Billiard | 2023-03-21 14:23:12 | View | |
29 Sep 2023
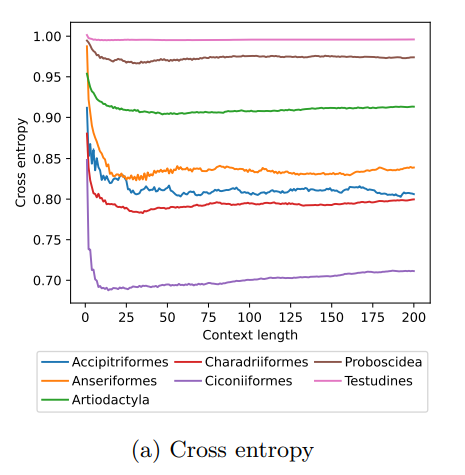
MoveFormer: a Transformer-based model for step-selection animal movement modellingOndřej Cífka, Simon Chamaillé-Jammes, Antoine Liutkus https://doi.org/10.1101/2023.03.05.531080A deep learning model to unlock secrets of animal movement and behaviourRecommended by Cédric Sueur based on reviews by Jacob Davidson and 1 anonymous reviewer based on reviews by Jacob Davidson and 1 anonymous reviewer
The study of animal movement is essential for understanding their behaviour and how ecological or global changes impact their routines [1]. Recent technological advancements have improved the collection of movement data [2], but limited statistical tools have hindered the analysis of such data [3–5]. Animal movement is influenced not only by environmental factors but also by internal knowledge and memory, which are challenging to observe directly [6,7]. Routine movement behaviours and the incorporation of memory into models remain understudied. Researchers have developed ‘MoveFormer’ [8], a deep learning-based model that predicts future movements based on past context, addressing these challenges and offering insights into the importance of different context lengths and information types. The model has been applied to a dataset of over 1,550 trajectories from various species, and the authors have made the MoveFormer source code available for further research. Inspired by the step-selection framework and efforts to quantify uncertainty in movement predictions, MoveFormer leverages deep learning, specifically the Transformer architecture, to encode trajectories and understand how past movements influence current and future ones – a critical question in movement ecology. The results indicate that integrating information from a few days to two or three weeks before the movement enhances predictions. The model also accounts for environmental predictors and offers insights into the factors influencing animal movements. Its potential impact extends to conservation, comparative analyses, and the generalisation of uncertainty-handling methods beyond ecology, with open-source code fostering collaboration and innovation in various scientific domains. Indeed, this method could be applied to analyse other kinds of movements, such as arm movements during tool use [9], pen movements, or eye movements during drawing [10], to better understand anticipation in actions and their intentionality. References 1. Méndez, V.; Campos, D.; Bartumeus, F. Stochastic Foundations in Movement Ecology: Anomalous Diffusion, Front Propagation and Random Searches; Springer Series in Synergetics; Springer: Berlin, Heidelberg, 2014; ISBN 978-3-642-39009-8. | MoveFormer: a Transformer-based model for step-selection animal movement modelling | Ondřej Cífka, Simon Chamaillé-Jammes, Antoine Liutkus | <p style="text-align: justify;">The movement of animals is a central component of their behavioural strategies. Statistical tools for movement data analysis, however, have long been limited, and in particular, unable to account for past movement i... |  | Behaviour & Ethology, Habitat selection | Cédric Sueur | 2023-03-22 16:32:14 | View | |
27 Nov 2023
Modeling Tick Populations: An Ecological Test Case for Gradient Boosted TreesWilliam Manley, Tam Tran, Melissa Prusinski, Dustin Brisson https://doi.org/10.1101/2023.03.13.532443Gradient Boosted Trees can deliver more than accurate ecological predictionsRecommended by Timothée Poisot based on reviews by 2 anonymous reviewers based on reviews by 2 anonymous reviewers
Tick-borne diseases are an important burden on public health all over the globe, making accurate forecasts of tick population a key ingredient in a successful public health strategy. Over long time scales, tick populations can undergo complex dynamics, as they are sensitive to many non-linear effects due to the complex relationships between ticks and the relevant (numerical) features of their environment. But luckily, capturing complex non-linear responses is a task that machine learning thrives on. In this contribution, Manley et al. (2023) explore the use of Gradient Boosted Trees to predict the distribution (presence/absence) and abundance of ticks across New York state. This is an interesting modelling challenge in and of itself, as it looks at the same ecological question as an instance of a classification problem (presence/absence) or of a regression problem (abundance). In using the same family of algorithm for both, Manley et al. (2023) provide an interesting showcase of the versatility of these techniques. But their article goes one step further, by setting up a multi-class categorical model that estimates jointly the presence and abundance of a population. I found this part of the article particularly elegant, as it provides an intermediate modelling strategy, in between having two disconnected models for distribution and abundance, and having nested models where abundance is only predicted for the present class (see e.g. Boulangeat et al., 2012, for a great description of the later). One thing that Manley et al. (2023) should be commended for is their focus on opening up the black box of machine learning techniques. I have never believed that ML models are more inherently opaque than other families of models, but the focus in this article on explainable machine learning shows how these models might, in fact, bring us closer to a phenomenological understanding of the mechanisms underpinning our observations. There is also an interesting discussion in this article, on the rate of false negatives in the different models that are being benchmarked. Although model selection often comes down to optimizing the overall quality of the confusion matrix (for distribution models, anyway), depending on the type of information we seek to extract from the model, not all types of errors are created equal. If the purpose of the model is to guide actions to control vectors of human pathogens, a false negative (predicting that the vector is absent at a site where it is actually present) is a potentially more damaging outcome, as it can lead to the vector population (and therefore, potentially, transmission) increasing unchecked. References
Boulangeat I, Gravel D, Thuiller W. Accounting for dispersal and biotic interactions to disentangle the drivers of species distributions and their abundances: The role of dispersal and biotic interactions in explaining species distributions and abundances. Ecol Lett. 2012;15: 584-593. Manley W, Tran T, Prusinski M, Brisson D. (2023) Modeling tick populations: An ecological test case for gradient boosted trees. bioRxiv, 2023.03.13.532443, ver. 3 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2023.03.13.532443 | Modeling Tick Populations: An Ecological Test Case for Gradient Boosted Trees | William Manley, Tam Tran, Melissa Prusinski, Dustin Brisson | <p style="text-align: justify;">General linear models have been the foundational statistical framework used to discover the ecological processes that explain the distribution and abundance of natural populations. Analyses of the rapidly expanding ... | Parasitology, Species distributions, Statistical ecology | Timothée Poisot | Anonymous, Anonymous | 2023-03-23 23:41:17 | View | |
10 Aug 2023
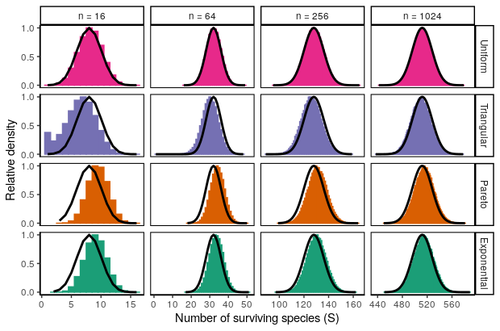
Coexistence of many species under a random competition-colonization trade-offZachary R. Miller, Maxime Clenet, Katja Della Libera, François Massol, Stefano Allesina https://doi.org/10.1101/2023.03.23.533867Assembly in metacommunities driven by a competition-colonization tradeoff: more species in, more species outRecommended by Frederik De Laender based on reviews by Canan Karakoç and 1 anonymous reviewerThe output of a community model depends on how you set its parameters. Thus, analyses of specific parameter settings hardwire the results to specific ecological scenarios. Because more general answers are often of interest, one tradition is to give models a statistical treatment: one summarizes how model parameters vary across species, and then predicts how changing the summary, instead of the individual parameters themselves, would change model output. Arguably the best-known example is the work initiated by May, showing that the properties of a community matrix, encoding effects species have on each other near their equilibrium, determine stability (1,2). More recently, this statistical treatment has also been applied to one of community ecology’s more prickly and slippery subjects: community assembly, which deals with the question “Given some regional species pool, which species will be able to persist together at some local ecosystem?”. Summaries of how species grow and interact in this regional pool predict the fraction of survivors and their relative abundances, the kind of dynamics, and various kinds of stability (3,4). One common characteristic of such statistical treatments is the assumption of disorder: if species do not interact in too structured ways, simple and therefore powerful predictions ensue that often stand up to scrutiny in relatively ordered systems. 2. Allesina, S. & Tang, S. (2015). The stability–complexity relationship at age 40: a random matrix perspective. Population Ecology, 57, 63–75. https://doi.org/10.1007/s10144-014-0471-0 3. Bunin, G. (2016). Interaction patterns and diversity in assembled ecological communities. Preprint at http://arxiv.org/abs/1607.04734. 5. Miller, Z. R., Clenet, M., Libera, K. D., Massol, F. & Allesina, S. (2023). Coexistence of many species under a random competition-colonization trade-off. bioRxiv 2023.03.23.533867, ver 3 peer-reviewed and recommended by PCI Ecology. https://doi.org/10.1101/2023.03.23.533867 6. Serván, C. A. & Allesina, S. (2021). Tractable models of ecological assembly. Ecology Letters, 24, 1029–1037. https://doi.org/10.1111/ele.13702 | Coexistence of many species under a random competition-colonization trade-off | Zachary R. Miller, Maxime Clenet, Katja Della Libera, François Massol, Stefano Allesina | <p>The competition-colonization trade-off is a well-studied coexistence mechanism for metacommunities. In this setting, it is believed that coexistence of all species requires their traits to satisfy restrictive conditions limiting their similarit... |  | Biodiversity, Coexistence, Colonization, Community ecology, Competition, Population ecology, Spatial ecology, Metacommunities & Metapopulations, Theoretical ecology | Frederik De Laender | 2023-03-30 20:42:48 | View | |
07 Aug 2023
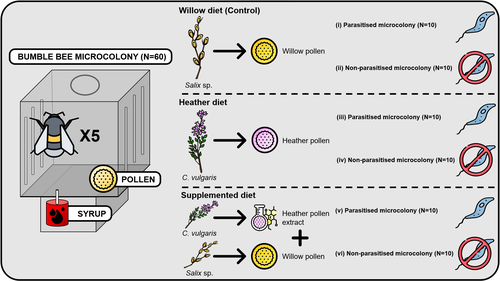
Heather pollen is not necessarily a healthy diet for bumble beesClément Tourbez, Irène Semay, Apolline Michel, Denis Michez, Pascal Gerbaux, Antoine Gekière, Maryse Vanderplanck https://doi.org/10.5281/zenodo.8192036The importance of understanding bee nutritionRecommended by Ignasi Bartomeus based on reviews by Cristina Botías and 1 anonymous reviewer based on reviews by Cristina Botías and 1 anonymous reviewer
Contrasting with the great alarm on bee declines, it is astonishing how little basic biology we know about bees, including on abundant and widespread species that are becoming model species. Plant-pollinator relationships are one of the cornerstones of bee ecology, and researchers are increasingly documenting bees' diets. However, we rarely know which effects feeding on different flowers has on bees' health. This paper (Tourbez et al. 2023) uses an elegant experimental setting to test the effect of heather pollen on bumblebees' (Bombus terrestris) reproductive success. This is a timely question as heather is frequently used by bumblebees, and its nectar has been reported to reduce parasite infections. In fact, it has been suggested that bumblebees can medicate themselves when infected (Richardson et al. 2014), and the pollen of some Asteraceae has been shown to help them fight parasites (Gekière et al. 2022). The starting hypothesis is that heather pollen contains flavonoids that might have a similar effect. Unfortunately, Tourbez and collaborators do not support this hypothesis, showing a negative effect of heather pollen, in particular its flavonoids, in bumblebees offspring, and an increase in parasite loads when fed on flavonoids. This is important because it challenges the idea that many pollen and nectar chemical compounds might have a medicinal use, and force us to critically analyze the effect of chemical compounds in each particular case. The results open several questions, such as why bumblebees collect heather pollen, or in which concentrations or pollen mixes it is deleterious. A limitation of the study is that it uses micro-colonies, and extrapolating this to real-world conditions is always complex. Understanding bee declines require a holistic approach starting with bee physiology and scaling up to multispecies population dynamics. References Gekière, A., Semay, I., Gérard, M., Michez, D., Gerbaux, P., & Vanderplanck, M. 2022. Poison or Potion: Effects of Sunflower Phenolamides on Bumble Bees and Their Gut Parasite. Biology, 11(4), 545. https://doi.org/10.3390/biology11040545 Richardson, L.L., Adler, L.S., Leonard, A.S., Andicoechea, J., Regan, K.H., Anthony, W.E., Manson, J.S., & Irwin, R.E. 2015. Secondary metabolites in floral nectar reduce parasite infections in bumblebees. Proceedings of the Royal Society of London B: Biological Sciences 282 (1803), 20142471. https://doi.org/10.1098/rspb.2014.2471 Tourbez, C., Semay, I., Michel, A., Michez, D., Gerbaux, P., Gekière A. & Vanderplanck, M. 2023. Heather pollen is not necessarily a healthy diet for bumble bees. Zenodo, ver 3, reviewed and recommended by PCI Ecology. https://doi.org/10.5281/zenodo.8192036 | Heather pollen is not necessarily a healthy diet for bumble bees | Clément Tourbez, Irène Semay, Apolline Michel, Denis Michez, Pascal Gerbaux, Antoine Gekière, Maryse Vanderplanck | <p>There is evidence that specialised metabolites of flowering plants occur in both vegetative parts and floral resources (i.e., pollen and nectar), exposing pollinators to their biological activities. While such metabolites may be toxic to bees, ... |  | Botany, Chemical ecology, Host-parasite interactions, Pollination, Zoology | Ignasi Bartomeus | 2023-04-10 21:22:34 | View | |
28 Aug 2023
Implementing a rapid geographic range expansion - the role of behavior changesLogan CJ, McCune KB, LeGrande-Rolls C, Marfori Z, Hubbard J, Lukas D https://doi.org/10.32942/X2N30JBehavioral changes in the rapid geographic expansion of the great-tailed grackleRecommended by Esther Sebastián González based on reviews by Francois-Xavier Dechaume-Moncharmont, Pizza Ka Yee Chow and 1 anonymous reviewer based on reviews by Francois-Xavier Dechaume-Moncharmont, Pizza Ka Yee Chow and 1 anonymous reviewer
While many species' populations are declining, primarily due to human-related impacts (McKnee et al., 2014), certain species have thrived by utilizing human-influenced environments, leading to their population expansion (Muñoz & Real, 2006). In this context, the capacity to adapt and modify behaviors in response to new surroundings is believed to play a crucial role in facilitating species' spread to novel areas (Duckworth & Badyaev, 2007). For example, an increase in innovative behaviors within recently established communities could aid in discovering previously untapped food resources, while a decrease in exploration might reduce the likelihood of encountering dangers in unfamiliar territories (e.g., Griffin et al., 2016). To investigate the contribution of these behaviors to rapid range expansions, it is essential to directly measure and compare behaviors in various populations of the species. The study conducted by Logan et al. (2023) aims to comprehend the role of behavioral changes in the range expansion of great-tailed grackles (Quiscalus mexicanus). To achieve this, the researchers compared the prevalence of specific behaviors at both the expansion's edge and its middle. Great-tailed grackles were chosen as an excellent model due to their behavioral adaptability, rapid geographic expansion, and their association with human-modified environments. The authors carried out a series of experiments in captivity using wild-caught individuals, following a detailed protocol. The study successfully identified differences in two of the studied behavioral traits: persistence (individuals participated in a larger proportion of trials) and flexibility variance (a component of the species' behavioral flexibility, indicating a higher chance that at least some individuals in the population could be more flexible). Notably, individuals at the edge of the population exhibited higher values of persistence and flexibility, suggesting that these behavioral traits might be contributing factors to the species' expansion. Overall, the study by Logan et al. (2023) is an excellent example of the importance of behavioral flexibility and other related behaviors in the process of species' range expansion and the significance of studying these behaviors across different populations to gain a better understanding of their role in the expansion process. Finally, it is important to underline that this study is part of a pre-registration that received an In Principle Recommendation in PCI Ecology (Sebastián-González 2020) where objectives, methodology, and expected results were described in detail. The authors have identified any deviation from the original pre-registration and thoroughly explained the reasons for their deviations, which were very clear. References Duckworth, R. A., & Badyaev, A. V. (2007). Coupling of dispersal and aggression facilitates the rapid range expansion of a passerine bird. Proceedings of the National Academy of Sciences, 104(38), 15017-15022. https://doi.org/10.1073/pnas.0706174104 Griffin, A.S., Guez, D., Federspiel, I., Diquelou, M., Lermite, F. (2016). Invading new environments: A mechanistic framework linking motor diversity and cognition to establishment success. Biological Invasions and Animal Behaviour, 26e46. https://doi.org/10.1017/CBO9781139939492.004 Logan, C. J., McCune, K., LeGrande-Rolls, C., Marfori, Z., Hubbard, J., Lukas, D. 2023. Implementing a rapid geographic range expansion - the role of behavior changes. EcoEvoRxiv, ver. 3 peer-reviewed and recommended by PCI Ecology. https://doi.org/10.32942/X2N30J McKee, J. K., Sciulli, P. W., Fooce, C. D., & Waite, T. A. (2004). Forecasting global biodiversity threats associated with human population growth. Biological Conservation, 115(1), 161-164. https://doi.org/10.1016/S0006-3207(03)00099-5 Muñoz, A. R., & Real, R. (2006). Assessing the potential range expansion of the exotic monk parakeet in Spain. Diversity and Distributions, 12(6), 656-665. https://doi.org/10.1111/j.1472-4642.2006.00272.x Sebastián González, E. (2020) The role of behavior and habitat availability on species geographic expansion. Peer Community in Ecology, 100062. https://doi.org/10.24072/pci.ecology.100062. Reviewers: Caroline Nieberding, Tim Parker, and Pizza Ka Yee Chow. | Implementing a rapid geographic range expansion - the role of behavior changes | Logan CJ, McCune KB, LeGrande-Rolls C, Marfori Z, Hubbard J, Lukas D | <p>It is generally thought that behavioral flexibility, the ability to change behavior when circumstances change, plays an important role in the ability of species to rapidly expand their geographic range. Great-tailed grackles (<em>Quiscalus mexi... | Behaviour & Ethology, Preregistrations, Zoology | Esther Sebastián González | 2023-04-12 11:00:42 | View | ||
03 Jan 2024
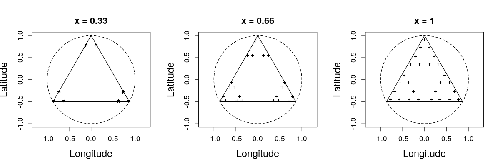
Efficient sampling designs to assess biodiversity spatial autocorrelation : should we go fractal?Fabien Laroche https://doi.org/10.1101/2022.07.29.501974Spatial patterns and autocorrelation challenges in ecological conservationRecommended by Eric Goberville based on reviews by Nigel Yoccoz and Charles J Marsh based on reviews by Nigel Yoccoz and Charles J Marsh
“Pattern, like beauty, is to some extent in the eye of the beholder” (Grant 1977 in Wiens, 1989) Ecologists are immersed in unraveling the complex spatial patterns that govern species diversity, driven by both practical and theoretical imperatives (Rahbek, 2005; Wang et al., 2019). This dual focus necessitates a practical imperative for strategic biodiversity conservation, requiring a nuanced understanding of locations with peak species richness and dynamic shifts in species assemblages (Chase et al., 2020). Simultaneously, there is a theoretical interest in using diversity patterns as empirical testing grounds for theories explaining factors influencing diversity disparities and the associated increase in species turnover correlated with inter-site distance (Condit et al., 2002).
McGill (2010), in his paper "Matters of Scale", highlights the scale-dependent nature of ecology, aligning with the recognition that spatial autocorrelation is inherent in biogeographical data and often correlated with sample size (Rahbek, 2005). Spatial autocorrelation, often underestimated in ecological studies (Dormann, 2007), occurs when proximate locations exhibit similarities in ecological attributes (Tobler, 1970; Getis, 2010), introducing a latent bias that compromises the robustness of ecological findings (Dormann, 2007; Dormann et al., 2007). This phenomenon serves as both an asset, providing valuable information for inferring processes from patterns (Palma et al. 1999), and a challenge, imposing limitations on hypothesis testing and prediction (Dormann et al., 2007 and references therein). Various factors contribute to spatial autocorrelation, with three primary contributors (Dormann et al., 2007; Legendre, 1993; Legendre and Fortin, 1989; Legendre and Legendre, 2012): (i) distance-related effects in biological processes, (ii) misrepresentation of non-linear relationships between the environment and species as linear and (iii) the oversight of a crucial spatially structured environmental determinant in the statistical model, leading to spatial structuring in the response (Dormann et al., 2007).
Recognising the pivotal role of spatial heterogeneity in ecological theories (Wang et al., 2019), it becomes imperative to discern and address the limitations introduced by spatial autocorrelation (Legendre, 1993). McGill (2011) emphasises that the ultimate goal of biodiversity pattern studies should be to develop a quantitative predictive theory useful for conservation. The spatial dimension's importance in study planning, determining the system's scale, appropriate quadrat size, and spacing between sampling stations, is paramount (Fortin, 1999a,b). Responses to these considerations are intricately linked with study objectives and insights from pre-sampling campaigns, underscoring the need for a nuanced and rigorous approach (Delmelle, 2021).
Understanding statistical techniques and nested sampling designs is crucial to answering fundamental ecological questions (Dormann et al., 2007; McDonald, 2012). In addressing spatial autocorrelation challenges, ecologists must recognize the limitations of many standard statistical methods in ecological studies (Dale and Fortin, 2002; Legendre and Fortin, 1989; Steel et al., 2013). In the initial phases of description or hypothesis generation, ecologists should proactively acknowledge the spatial structure in their data and conduct tests for spatial autocorrelation (for a comprehensive description, see Legendre and Fortin, 1989): various tools, including correlograms, spectral analysis, the Mantel test, and clustering methods, facilitate the assessment and description of spatial structures. The partial Mantel test enables the study of causal models with space as an explanatory variable. Techniques for mapping ecological variables, such as interpolation, trend surface analysis, and constrained clustering, yield maps providing valuable insights into the spatial dynamics of ecological systems.
This refined consideration of spatial autocorrelation emerges as an imperative in ecological research, fostering a deeper and more precise understanding of the intricate interplay between species diversity, spatial patterns, and the inherent limitations imposed by spatial autocorrelation (Legendre et al., 2002). This not only contributes significantly to the scientific discourse in ecology but also aligns with McGill's vision of developing predictive theories for effective conservation (Bacaro et al., 2016; McGill, 2011).
In this study by Fabien Laroche (2023), titled “Efficient sampling designs to assess biodiversity spatial autocorrelation: should we go fractal?” the primary focus was on addressing the challenges associated with estimating the autocorrelation range of species distribution across spatial scales. The study aimed to explore alternative sampling designs, with a particular focus on the application of fractal designs—self-similar designs with well-identified scales. The overarching goal was to evaluate whether fractal designs could offer a more efficient compromise compared to traditional hybrid designs, which involve mixing random sampling points with a systematic grid.
Virtual ecology provides a way to test whether sampling designs can accurately detect or quantify effects of interest before implementing them in the field. Beyond the question of assessing the power of empirical designs, a virtual ecology analysis contributes to clearly formulating the set of questions associated with a design. However, only a few virtual studies have focused on efficient designs to accurately estimate the autocorrelation range of biodiversity variables. In this study, the statistical framework of optimal design of experiments was employed—a methodology often used in building and comparing designs of temporal or spatiotemporal biodiversity surveys but rarely applied to the specific problem of quantifying spatial autocorrelation.
Key findings from the study shed light on optimal sampling strategies, with a notable dependence on the feasible grid mesh size over the study area in relation to expected autocorrelation range values. The results demonstrated that the efficiency of designs varied based on the specific effect under study. Fractal designs, however, exhibited superior performance, particularly when assessing the effect of a monotonic environmental gradient across space.
In conclusion, the study provides valuable insights into the potential benefits of incorporating fractal designs in biodiversity studies, offering a nuanced and efficient approach to estimate spatial autocorrelation. These findings contribute significantly to the ongoing scientific discourse in ecology, providing practical considerations for improving sampling designs in biodiversity assessments.
References
Bacaro, G., Altobelli, A., Cameletti, M., Ciccarelli, D., Martellos, S., Palmer, M.W., Ricotta, C., Rocchini, D., Scheiner, S.M., Tordoni, E., Chiarucci, A., 2016. Incorporating spatial autocorrelation in rarefaction methods: Implications for ecologists and conservation biologists. Ecological Indicators 69, 233-238. https://doi.org/10.1016/j.ecolind.2016.04.026
Chase, J.M., Jeliazkov, A., Ladouceur, E., Viana, D.S., 2020. Biodiversity conservation through the lens of metacommunity ecology. Annals of the New York Academy of Sciences 1469, 86-104. https://doi.org/10.1111/nyas.14378
Condit, R., Pitman, N., Leigh, E.G., Chave, J., Terborgh, J., Foster, R.B., Núñez, P., Aguilar, S., Valencia, R., Villa, G., Muller-Landau, H.C., Losos, E., Hubbell, S.P., 2002. Beta-Diversity in Tropical Forest Trees. Science 295, 666-669. https://doi.org/10.1126/science.1066854
Dale, M.R.T., Fortin, M.-J., 2002. Spatial autocorrelation and statistical tests in ecology. Écoscience 9, 162-167. https://doi.org/10.1080/11956860.2002.11682702
Delmelle, E.M., 2021. Spatial Sampling, in: Fischer, M.M., Nijkamp, P. (Eds.), Handbook of Regional Science. Springer Berlin Heidelberg, Berlin, Heidelberg, pp. 1829-1844.
Dormann, C.F., 2007. Effects of incorporating spatial autocorrelation into the analysis of species distribution data. Global Ecology & Biogeography 16, 129-128. https://doi.org/10.1111/j.1466-8238.2006.00279.x
Dormann, C.F., McPherson, J.M., Araújo, M.B., Bivand, R., Bolliger, J., Carl, G., Davies, R.G., Hirzel, A., Jetz, W., Kissling, W.D., Kühn, I., Ohlemüler, R., Peres-Neto, P.R., Reineking, B., Schröder, B., Schurr, F.M., Wilson, R., 2007. Methods to account for spatial autocorrelation in the analysis of species distributional data: a review. Ecography 33, 609-628. https://doi.org/10.1111/j.2007.0906-7590.05171.x
Fortin, M.-J., 1999a. Effects of quadrat size and data measurement on the detection of boundaries. Journal of Vegetation Science 10, 43-50. https://doi.org/10.2307/3237159
Fortin, M.-J., 1999b. Effects of sampling unit resolution on the estimation of spatial autocorrelation. Écoscience 6, 636-641. https://doi.org/10.1080/11956860.1999.11682547
Getis, A., 2010. Spatial Autocorrelation, in: Fischer, M.M., Getis, A. (Eds.), Handbook of Applied Spatial Analysis: Software Tools, Methods and Applications. Springer Berlin Heidelberg, Berlin, Heidelberg, pp. 255-278.
Laroche, F., 2023. Efficient sampling designs to assess biodiversity spatial autocorrelation: should we go fractal? bioRxiv, 2022.07.29.501974, ver. 4 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2022.07.29.501974
Legendre, P., 1993. Spatial Autocorrelation: Trouble or New Paradigm? Ecology 74, 1659-1673. https://doi.org/10.2307/1939924
Legendre, P., Dale, M.R.T., Fortin, M.-J., Gurevitch, J., Hohn, M., Myers, D., 2002. The consequences of spatial structure for the design and analysis of ecological field surveys. Ecography 25, 601-615. https://doi.org/10.1034/j.1600-0587.2002.250508.x
Legendre, P., Fortin, M.J., 1989. Spatial pattern and ecological analysis. Vegetatio 80, 107-138. https://doi.org/10.1007/BF00048036
Legendre, P., Legendre, L., 2012. Numerical Ecology, Third Edition ed. Elsevier, The Netherlands.
McDonald, T., 2012. Spatial sampling designs for long-term ecological monitoring, in: Cooper, A.B., Gitzen, R.A., Licht, D.S., Millspaugh, J.J. (Eds.), Design and Analysis of Long-term Ecological Monitoring Studies. Cambridge University Press, Cambridge, pp. 101-125.
McGill, B.J., 2010. Matters of Scale. Science 328, 575-576. https://doi.org/10.1126/science.1188528
McGill, B.J., 2011. Linking biodiversity patterns by autocorrelated random sampling. American Journal of Botany 98, 481-502. https://doi.org/10.3732/ajb.1000509
Rahbek, C., 2005. The role of spatial scale and the perception of large-scale species-richness patterns. Ecology Letters 8, 224-239. https://doi.org/10.1111/j.1461-0248.2004.00701.x
Steel, E.A., Kennedy, M.C., Cunningham, P.G., Stanovick, J.S., 2013. Applied statistics in ecology: common pitfalls and simple solutions. Ecosphere 4, art115. https://doi.org/10.1890/ES13-00160.1
Tobler, W.R., 1970. A Computer Movie Simulating Urban Growth in the Detroit Region. Economic Geography 46, 234-240. https://doi.org/10.2307/143141
Wang, S., Lamy, T., Hallett, L.M., Loreau, M., 2019. Stability and synchrony across ecological hierarchies in heterogeneous metacommunities: linking theory to data. Ecography 42, 1200-1211. https://doi.org/10.1111/ecog.04290
Wiens, J.A., 1989. The ecology of bird communities. Cambridge University Press.
| Efficient sampling designs to assess biodiversity spatial autocorrelation : should we go fractal? | Fabien Laroche | <p>Quantifying the autocorrelation range of species distribution in space is necessary for applied ecological questions, like implementing protected area networks or monitoring programs. However, the power of spatial sampling designs to estimate t... |  | Biodiversity, Landscape ecology, Spatial ecology, Metacommunities & Metapopulations, Statistical ecology | Eric Goberville | 2023-04-21 10:54:29 | View | |
07 Aug 2023
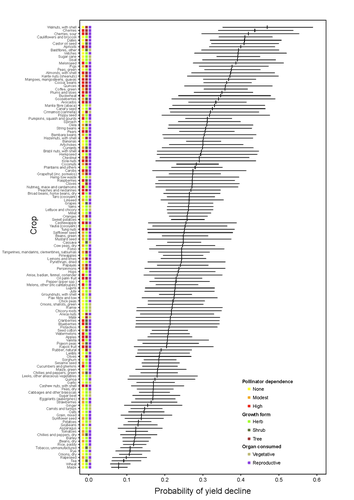
Being a tree crop increases the odds of experiencing yield declines irrespective of pollinator dependenceMarcelo A. Aizen, Gabriela Gleiser, Thomas Kitzberger, and Rubén Milla https://doi.org/10.1101/2023.04.27.538617The complexities of understanding why yield is decliningRecommended by Ignasi Bartomeus based on reviews by Nicolas Deguines and 1 anonymous reviewer based on reviews by Nicolas Deguines and 1 anonymous reviewer
Despite the repeated mantra that "correlation does not imply causation", ecological studies not amenable to experimental settings often rely on correlational patterns to infer the causes of observed patterns. In this context, it's of paramount importance to build a plausible hypothesis and take into account potential confounding factors. The paper by Aizen and collaborators (2023) is a beautiful example of how properly unveil the complexities of an intriguing pattern: The decline in yield of some crops over the last few decades. This is an outstanding question to solve given the need to feed a growing population without destroying the environment, for example by increasing the area under cultivation. Previous studies suggested that pollinator-dependent crops were more susceptible to suffering yield declines than non-pollinator-dependent crops (Garibaldi et al 2011). Given the actual population declines of some pollinators, especially in agricultural areas, this correlative evidence was quite appealing to be interpreted as a causal effect. However, as elegantly shown by Aizen and colleagues in this paper, this first analysis did not account for other alternative explanations, such as the effect of climate change on other plant life-history traits correlated with pollinator dependence. Plant life-history traits do not vary independently. For example, trees are more likely to be pollinator-dependent than herbs (Lanuza et al 2023), which can be an important confounding factor in the analysis. With an elegant analysis and an impressive global dataset, this paper shows that the declining trend in the yield of some crops is most likely associated with their life form than with their dependence on pollinators. This does not imply that pollinators are not important for crop yield, but that the decline in their populations is not leaving a clear imprint in the global yield production trends once accounted for the technological and agronomic improvements. All in all, this paper makes a key contribution to food security by elucidating the factors beyond declining yield trends, and is a brave example of how science can self-correct itself as new knowledge emerges. References Aizen, M.A., Gleiser, G., Kitzberger T. and Milla R. 2023. Being A Tree Crop Increases the Odds of Experiencing Yield Declines Irrespective of Pollinator Dependence. bioRxiv, 2023.04.27.538617, ver 2, peer-reviewed and recommended by PCI Ecology. https://doi.org/10.1101/2023.04.27.538617 Lanuza, J.B., Rader, R., Stavert, J., Kendall, L.K., Saunders, M.E. and Bartomeus, I. 2023. Covariation among reproductive traits in flowering plants shapes their interactions with pollinators. Functional Ecology 37: 2072-2084. https://doi.org/10.1111/1365-2435.14340 Garibaldi, L.A., Aizen, M.A., Klein, A.M., Cunningham, S.A. and Harder, L.D. 2011. Global growth and stability of agricultural yield decrease with pollinator dependence. Proceedings of the National Academy of Sciences, 108: 5909-5914. https://doi.org/10.1073/pnas.1012431108 | Being a tree crop increases the odds of experiencing yield declines irrespective of pollinator dependence | Marcelo A. Aizen, Gabriela Gleiser, Thomas Kitzberger, and Rubén Milla | <p>Crop yields, i.e., harvestable production per unit of cropland area, are in decline for a number of crops and regions, but the drivers of this process are poorly known. Global decreases in pollinator abundance and diversity have been proposed a... |  | Agroecology, Climate change, Community ecology, Demography, Facilitation & Mutualism, Life history, Phenotypic plasticity, Pollination, Terrestrial ecology | Ignasi Bartomeus | 2023-05-02 18:54:44 | View | |
03 Oct 2023
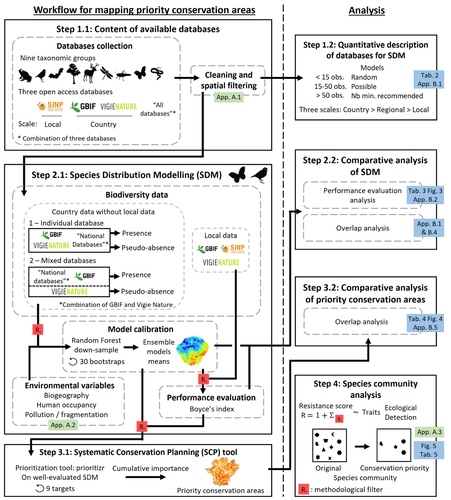
Integrating biodiversity assessments into local conservation planning: the importance of assessing suitable data sourcesThibaut Ferraille, Christian Kerbiriou, Charlotte Bigard, Fabien Claireau, John D. Thompson https://doi.org/10.1101/2023.05.09.539999Biodiversity databases are ever more numerous, but can they be used reliably for Species Distribution Modelling?Recommended by Nicolas Schtickzelle based on reviews by 2 anonymous reviewers based on reviews by 2 anonymous reviewers
Proposing efficient guidelines for biodiversity conservation often requires the use of forecasting tools. Species Distribution Models (SDM) are more and more used to predict how the distribution of a species will react to environmental change, including any large-scale management actions that could be implemented. Their use is also boosted by the increase of publicly available biodiversity databases[1]. The now famous aphorism by George Box "All models are wrong but some are useful"[2] very well summarizes that the outcome of a model must be adjusted to, and will depend on, the data that are used to parameterize it. The question of the reliability of using biodiversity databases to parameterize biodiversity models such as SDM –but the question would also apply to other kinds of biodiversity models, e.g. Population Viability Analysis models[3]– is key to determine the confidence that can be placed in model predictions. This point is often overlooked by some categories of biodiversity conservation stakeholders, in particular the fact that some data were collected using controlled protocols while others are opportunistic. In this study[4], the authors use a collection of databases covering a range of species as well as of geographic scales in France and using different data collection and validation approaches as a case study to evaluate the impact of data quality when performing Strategic Environmental Assessment (SEA). Among their conclusions, the fact that a large-scale database (what they call the “country” level) is necessary to reliably parameterize SDM. Besides this and other conclusions of their study, which are likely to be in part specific to their case study –unfortunately for its conservation, biodiversity is complex and varies a lot–, the merit of this work lies in the approach used to test the impact of data on model predictions. References 1. Feng, X. et al. A review of the heterogeneous landscape of biodiversity databases: Opportunities and challenges for a synthesized biodiversity knowledge base. Global Ecology and Biogeography 31, 1242–1260 (2022). https://doi.org/10.1111/geb.13497 2. Box, G. E. P. Robustness in the Strategy of Scientific Model Building. in Robustness in Statistics (eds. Launer, R. L. & Wilkinson, G. N.) 201–236 (Academic Press, 1979). https://doi.org/10.1016/B978-0-12-438150-6.50018-2. 3. Beissinger, S. R. & McCullough, D. R. Population Viability Analysis. (The University of Chicago Press, 2002). 4. Ferraille, T., Kerbiriou, C., Bigard, C., Claireau, F. & Thompson, J. D. (2023) Integrating biodiversity assessments into local conservation planning: the importance of assessing suitable data sources. bioRxiv, ver. 3 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2023.05.09.539999 | Integrating biodiversity assessments into local conservation planning: the importance of assessing suitable data sources | Thibaut Ferraille, Christian Kerbiriou, Charlotte Bigard, Fabien Claireau, John D. Thompson | <p>Strategic Environmental Assessment (SEA) of land-use planning is a fundamental tool to minimize environmental impacts of artificialization. In this context, Systematic Conservation Planning (SCP) tools based on Species Distribution Models (SDM)... |  | Biodiversity, Conservation biology, Species distributions, Terrestrial ecology | Nicolas Schtickzelle | 2023-05-11 09:41:05 | View | |
11 Aug 2023

Implementing Code Review in the Scientific Workflow: Insights from Ecology and Evolutionary BiologyEdward Ivimey-Cook, Joel Pick, Kevin Bairos-Novak, Antica Culina, Elliot Gould, Matthew Grainger, Benjamin Marshall, David Moreau, Matthieu Paquet, Raphaël Royauté, Alfredo Sanchez-Tojar, Inês Silva, Saras Windecker https://doi.org/10.32942/X2CG64A handy “How to” review code for ecologists and evolutionary biologistsRecommended by Corina Logan based on reviews by Serena Caplins and 1 anonymous reviewer based on reviews by Serena Caplins and 1 anonymous reviewer
Ivimey Cook et al. (2023) provide a concise and useful “How to” review code for researchers in the fields of ecology and evolutionary biology, where the systematic review of code is not yet standard practice during the peer review of articles. Consequently, this article is full of tips for authors on how to make their code easier to review. This handy article applies not only to ecology and evolutionary biology, but to many fields that are learning how to make code more reproducible and shareable. Taking this step toward transparency is key to improving research rigor (Brito et al. 2020) and is a necessary step in helping make research trustable by the public (Rosman et al. 2022). References Brito, J. J., Li, J., Moore, J. H., Greene, C. S., Nogoy, N. A., Garmire, L. X., & Mangul, S. (2020). Recommendations to enhance rigor and reproducibility in biomedical research. GigaScience, 9(6), giaa056. https://doi.org/10.1093/gigascience/giaa056 Ivimey-Cook, E. R., Pick, J. L., Bairos-Novak, K., Culina, A., Gould, E., Grainger, M., Marshall, B., Moreau, D., Paquet, M., Royauté, R., Sanchez-Tojar, A., Silva, I., Windecker, S. (2023). Implementing Code Review in the Scientific Workflow: Insights from Ecology and Evolutionary Biology. EcoEvoRxiv, ver 5 peer-reviewed and recommended by Peer Community In Ecology. https://doi.org/10.32942/X2CG64 Rosman, T., Bosnjak, M., Silber, H., Koßmann, J., & Heycke, T. (2022). Open science and public trust in science: Results from two studies. Public Understanding of Science, 31(8), 1046-1062. https://doi.org/10.1177/09636625221100686 | Implementing Code Review in the Scientific Workflow: Insights from Ecology and Evolutionary Biology | Edward Ivimey-Cook, Joel Pick, Kevin Bairos-Novak, Antica Culina, Elliot Gould, Matthew Grainger, Benjamin Marshall, David Moreau, Matthieu Paquet, Raphaël Royauté, Alfredo Sanchez-Tojar, Inês Silva, Saras Windecker | <p>Code review increases reliability and improves reproducibility of research. As such, code review is an inevitable step in software development and is common in fields such as computer science. However, despite its importance, code review is not... |  | Meta-analyses, Statistical ecology | Corina Logan | 2023-05-19 15:54:01 | View |
MANAGING BOARD
Julia Astegiano
Tim Coulson
Anna Eklof
Dominique Gravel
François Massol
Ben Phillips
Cyrille Violle










