
A deep learning model to unlock secrets of animal movement and behaviour
 based on reviews by Jacob Davidson and 1 anonymous reviewer
based on reviews by Jacob Davidson and 1 anonymous reviewer
MoveFormer: a Transformer-based model for step-selection animal movement modelling
Abstract
Recommendation: posted 29 September 2023, validated 29 September 2023
Sueur, C. (2023) A deep learning model to unlock secrets of animal movement and behaviour. Peer Community in Ecology, 100531. https://doi.org/10.24072/pci.ecology.100531
Recommendation
The study of animal movement is essential for understanding their behaviour and how ecological or global changes impact their routines [1]. Recent technological advancements have improved the collection of movement data [2], but limited statistical tools have hindered the analysis of such data [3–5]. Animal movement is influenced not only by environmental factors but also by internal knowledge and memory, which are challenging to observe directly [6,7]. Routine movement behaviours and the incorporation of memory into models remain understudied.
Researchers have developed ‘MoveFormer’ [8], a deep learning-based model that predicts future movements based on past context, addressing these challenges and offering insights into the importance of different context lengths and information types. The model has been applied to a dataset of over 1,550 trajectories from various species, and the authors have made the MoveFormer source code available for further research.
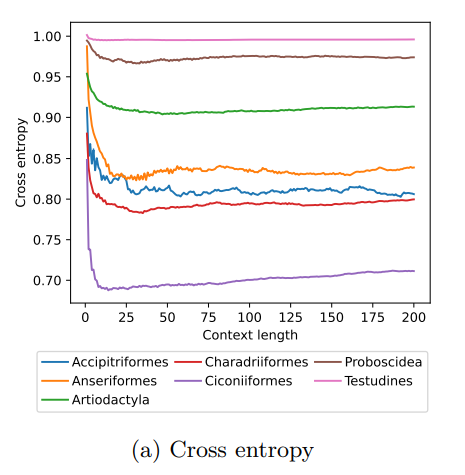
Inspired by the step-selection framework and efforts to quantify uncertainty in movement predictions, MoveFormer leverages deep learning, specifically the Transformer architecture, to encode trajectories and understand how past movements influence current and future ones – a critical question in movement ecology. The results indicate that integrating information from a few days to two or three weeks before the movement enhances predictions. The model also accounts for environmental predictors and offers insights into the factors influencing animal movements.
Its potential impact extends to conservation, comparative analyses, and the generalisation of uncertainty-handling methods beyond ecology, with open-source code fostering collaboration and innovation in various scientific domains. Indeed, this method could be applied to analyse other kinds of movements, such as arm movements during tool use [9], pen movements, or eye movements during drawing [10], to better understand anticipation in actions and their intentionality.
References
1. Méndez, V.; Campos, D.; Bartumeus, F. Stochastic Foundations in Movement Ecology: Anomalous Diffusion, Front Propagation and Random Searches; Springer Series in Synergetics; Springer: Berlin, Heidelberg, 2014; ISBN 978-3-642-39009-8.
https://doi.org/10.1007/978-3-642-39010-4
2. Fehlmann, G.; King, A.J. Bio-Logging. Curr. Biol. 2016, 26, R830-R831.
https://doi.org/10.1016/j.cub.2016.05.033
3. Jacoby, D.M.; Freeman, R. Emerging Network-Based Tools in Movement Ecology. Trends Ecol. Evol. 2016, 31, 301-314.
https://doi.org/10.1016/j.tree.2016.01.011
4. Michelot, T.; Langrock, R.; Patterson, T.A. moveHMM: An R Package for the Statistical Modelling of Animal Movement Data Using Hidden Markov Models. Methods Ecol. Evol. 2016, 7, 1308-1315.
https://doi.org/10.1111/2041-210X.12578
5. Wang, G. Machine Learning for Inferring Animal Behavior from Location and Movement Data. Ecol. Inform. 2019, 49, 69-76.
https://doi.org/10.1016/j.ecoinf.2018.12.002
6. Noser, R.; Byrne, R.W. Change Point Analysis of Travel Routes Reveals Novel Insights into Foraging Strategies and Cognitive Maps of Wild Baboons. Am. J. Primatol. 2014, 76, 399-409.
https://doi.org/10.1002/ajp.22181
7. Fagan, W.F.; Lewis, M.A.; Auger‐Méthé, M.; Avgar, T.; Benhamou, S.; Breed, G.; LaDage, L.; Schlägel, U.E.; Tang, W.; Papastamatiou, Y.P. Spatial Memory and Animal Movement. Ecol. Lett. 2013, 16, 1316-1329.
https://doi.org/10.1111/ele.12165
8. Cífka, O.; Chamaillé-Jammes, S.; Liutkus, A. MoveFormer: A Transformer-Based Model for Step-Selection Animal Movement Modelling. bioRxiv 2023, ver. 4 peer-reviewed and recommended by Peer Community in Ecology.
https://doi.org/10.1101/2023.03.05.531080
9. Ardoin, T.; Sueur, C. Automatic Identification of Stone-Handling Behaviour in Japanese Macaques Using LabGym Artificial Intelligence. 2023, https://doi.org/10.13140/RG.2.2.30465.02402
10. Martinet, L.; Pelé, M. Drawing in Nonhuman Primates: What We Know and What Remains to Be Investigated. J. Comp. Psychol. Wash. DC 1983 2021, 135, 176-184, doi:10.1037/com0000251.
https://doi.org/10.1037/com0000251
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
This work was supported by the LabEx NUMEV (ANR-10-LABX-0020) and the REPOS project, both funded by the I-Site MUSE (ANR-16-IDEX-0006). Computations were performed using HPC/AI resources from GENCI-IDRIS (Grant AD011012019R1).
Evaluation round #1
DOI or URL of the preprint: https://doi.org/10.1101/2023.03.05.531080
Version of the preprint: 2
Author's Reply, 27 Sep 2023
Dear recommender,
please see our response to reviews in the pdf. We also provide a version with track-change so you can more immediately see the revisions made. The document with these changes accepted is the lastest version online on bioRxiv.
We hope this revision will match your expectations.
Best regards,
Simon Chamaillé-Jammes, on behalf of the authors.
Decision by Cédric Sueur , posted 18 May 2023, validated 21 May 2023
, posted 18 May 2023, validated 21 May 2023
Dear authors,
We received the comments of two reviewers. Please prepare a response. We will be pleased to review another version of your manuscript.
All the best,
Cédric Sueur
Reviewed by Jacob Davidson, 17 May 2023
The authors develop a machine learning approach for predicting animal trajectories that uses the Transformer network architecture in order to incorporate past information of multiple features into predictions. By fitting to data from many different species using openly available MoveBank data, the authors compare predictive ability for different features and as a function of how much time of previous data is included.
I think the approach is interesting and makes a contribution that others working with movement data can use and build on. I have one main concern about the fitting procedure though: mainly, can the conclusions comparing the species be made, when the model is fit to all data together and the number of data points varies so large between species? The authors also note this (line 468). Does this discrepancy of data affect the conclusions? I could imagine an alternative fitting procedure, where each species is weighted equally, instead of each trajectory point. I feel that this comparison, or else further description justifying why the species comparison is driven by behavioral differences instead of simply different amounts of data, is needed.
Minor comments:
Legend text on Fig 1 is too small, and lacks units
Fig 2 shows PCA results for comparing the species, but does not show the PCA vectors. I'm not familiar with the Wikipedia2Vec data, but for PCA the vector components are normally shown, so that one can see what the embeddings represent. If this is not relevant for showing the Wikipedia2Vec embeddings, then it should at least be mentioned.
https://doi.org/10.24072/pci.ecology.100531.rev11Reviewed by anonymous reviewer 1, 12 May 2023
# General comments
In this paper, the authors propose a deep learning (neural network) model for analysing animal movement trajectories, called MoveFormer. The model is step-based, predicting an animal’s next step based on the environmental context (as in a step-selection function). However, the model learns the entire trajectory before that step, thus incorporating (potentially long) temporal context to make predictions. Being a deep learning approach, we expect that the model is capable of learning complex relationships and having high predictive power. I believe that a similar deep learning approach for analysing trajectories (sequences) is Long Short Term Memory, but the paper uses recent developments such as the Transformer architecture. In my reading, I have not seen a similar approach applied to trajectory data. In general, I think this is a useful contribution to movement ecology: I feel we will (and should) see more approaches like this, which leverage the potential of deep learning methods for incorporating sequence (temporal) information when analysing animal trajectories. Specifically, the incorporation of previous movements (history) is an important advantage of the approach, and the estimation of a ‘context length’ (time window) that is most important for being able to learn the trajectory is a key contribution.
I have some familiarity with simpler machine learning methods, but not much expertise in deep learning. I cannot comment on many technical aspects of the work, particularly the specific architecture and implementation of this model. Nonetheless, I offer below a few general comments, and a small number of specific comments.
Clearly the model has impressive predictive capability – I wondered whether it’s possible to forecast more than one step ahead, or to predict a whole sequence of the same output length as the input? I understand that this is no longer exactly step selection, then, but I think this would be the kind of application many movement ecologists would be interested in. If we can only predict one step ahead, then a simpler approach may be better (next point)?
I was curious about inference from the model. If the model cannot (or should not) be used to predict more than one step ahead, and inference is limited due to the black box nature of the model, is it better to sticks with more traditional step selection functions if inference is the goal? I guess this is particularly relevant given how much data are clearly required for the present model. Regarding inference, is it possible to look at ‘selection’ of environmental conditions, along with the importance currently shown in the paper?
I appreciated that some features of the model – I’m thinking particularly of the different time-scale periods in the model – were kept general to maximise the wider (future) application of the model.
Regarding the stated contributions number 2 and 3 -- “Second, the proposed approach is flexible enough to allow each step in the context to be defined not only by the locations of the start and end points, but also by any kind of features that could be relevant, in particular environmental variables. Third, we show how the model can be used to gain insights about the importance of the provided context, both in terms of the extent of the past that it is useful to know, and in terms of what kinds of information are most ecologically relevant to predict an animal’s movement” – it would be really interesting (in future work!) to see how the model responds to variation in the spatial scale and resolution of the environmental context variables.
I found the evaluation of the relevant context length very interesting. In future work it would be interesting to further examine patterns of context length among species, beyond what is presented here, and in different ecological settings.
The present study uses GPS data – I would be interested to hear the authors’ thoughts on how to deal with lower accuracy tracking data such as Argos.
An important positive element of the work is the open-source release of the software, although I have not had the opportunity to try it.
Overall, the manuscript is clearly written and neatly presented.
# A few specific comments
L70-73: I don't agree that step selection functions are *the* approach to analyse animal trajectories. My own feeling is that other methods such as Hidden Markov Models or regression of trajectory parameters against environmental covariates are *at least* equally common, if not more so. To avoid this statement (which I think is debatable), consider: "Step-selection function (SSF) models, which compare actual movement steps with realistic candidate ones, are routinely used to infer and quantify the effect of environmental variables, such as land cover or temperature, on animal trajectories".
L105-107: I assume the time-window is arbitrary, which was one of your criticisms of current methods for incorporating previous context ("familiarity") in step selection functions.
Table 1: Add column name abbreviations to the table caption. Describe 'Section' part of the table (training, validation, test) in the caption.
L150: '408 observations'. Also, translate this approximately to a real duration in days?
L150-153: How is the split assigned (what proportions)?
L156-158: What is the reason for doing this?
L159: I think the taxon vectors will need some further explanation. Is the vector approximating the taxonomic relationships? Okay, I see this information down on L171-172. I suggest moving this information (or something like it) up to the beginning of the paragraph, to immediately give readers the context. Although, I still wonder if the vector embedding is capturing the actual taxonomic relationship, or only the ‘semantic similarity’ [L164-165] (in other words, how similar given Wikipedia entries are). The PCA figure, for example, shows that the embedding is okay at class level, but not very good at order level (no clear clustering of orders within classes). Also, the Spearman correlation (= 0.68, L 170) is not great. Could there be a different way to embed taxonomy? In understand that there may not be, and this is a minor point.
L184: ‘resample’ rather than ‘sample’?
L189: From this dataset (and Figure 1), I gather that no marine trajectories are included. Were these explicitly excluded using a filter at some point?
L312. Perhaps start a new paragraph here.
Figure 7: The labels ‘bioclim’ are not informative.
L502-505: I’m not sure I completely understood this point, but it seems to be quite relevant give our wish to make inference from the model. So, currently you present no inference for features that were relevant in the learned trajectory, only for the next predicted step? How are the two related to one another?
L537-540: I wonder, also, whether using higher temporal resolution data would reveal a second peak in context length, indicative of nested scale-patterns in the trajectories (e.g., fine temporal context nested within a longer context).
https://doi.org/10.24072/pci.ecology.100531.rev12









