Direct submissions to PCI Ecology from bioRxiv.org are possible using the B2J service
Latest recommendations

| Id | Title * | Authors * | Abstract * | Picture * | Thematic fields * | Recommender | Reviewers▲ | Submission date | |
|---|---|---|---|---|---|---|---|---|---|
07 Jun 2023
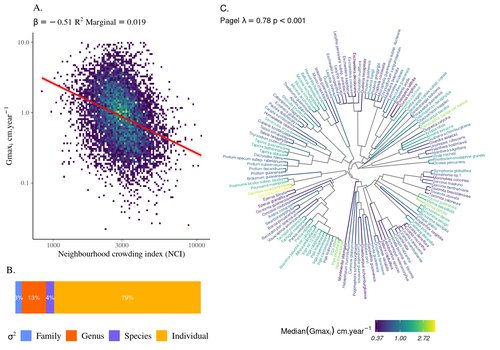
High intraspecific growth variability despite strong evolutionary heritage in a neotropical forestSylvain Schmitt, Bruno Hérault, Géraldine Derroire https://doi.org/10.1101/2022.07.27.501745Environmental and functional determinants of tree performance in a neotropical forest: the imprint of evolutionary legacy on growth strategiesRecommended by François Munoz based on reviews by David Murray-Stoker, Camille Girard and Jelena Pantel based on reviews by David Murray-Stoker, Camille Girard and Jelena Pantel
The hyperdiverse tropical forests have long fascinated ecologists because the fact that so many species persist at a low density at a local scale remains hard to explain. Both niche-based and neutral hypotheses have been tested, primarily based on analyzing the taxonomic composition of tropical forest plots (Janzen 1970; Hubbell 2001). Studies of the functional and phylogenetic structure of tropical tree communities have further aimed to better assess the importance of niche-based processes. For instance, Baraloto et al. (2012) found that co-occurring species were functionally and phylogenetically more similar in a neotropical forest, suggesting a role of environmental filtering. Likewise, Schmitt et al. (2021) found the influence of environmental filtering on the functional composition of an Indian rainforest. Yet these studies evidenced non-random trait-environment association based on the composition of assemblages only (in terms of occurrences and abundances). A major challenge remains to further address whether and how tree performance varies among species and individuals in tropical forests. Functional traits are related to components of individual fitness (Violle et al. 2007). Recently, more and more emphasis has been put on examining the relationship between functional trait values and demographic parameters (Salguero-Gómez et al. 2018), in order to better understand how functional trait values determine species population dynamics and abundances in assemblages. Fortunel et al. (2018) found an influence of functional traits on species growth variation related to topography, and less clearly to neighborhood density (crowding). Poorter et al. (2018) observed 44% of trait variation within species in a neotropical forest. Although individual trait values would be expected to be better predictors of performance than average values measured at the species level, Poorter et al still found a poor relationship. Schmitt et al. (2023) examined how abiotic conditions and biotic interactions (considering neighborhood density) influenced the variation of individual potential tree growth, in a tropical forest plot located in French Guiana. They also considered the link between species-averaged values of growth potential and functional traits. Schmitt et al. (2023) found substantial variation in growth potential within species, that functional traits explained 40% of the variation of species-averaged growth and, noticeably, that the taxonomic structure (used as random effect in their model) explained a third of the variation in individual growth. Although functional traits of roots, wood and leaves could predict a significant part of species growth potential, much variability of tree growth occurred within species. Intraspecific trait variation can thus be huge in response to changing abiotic and biotic contexts across individuals. The information on phylogenetic relationships can still provide a proxy of the integrated phenotypic variation that is under selection across the phylogeny, and determine a variation in growth strategies among individuals. The similarity of the phylogenetic structure suggests a joint selection of these growth strategies and related functional traits during events of convergent evolution. Baraloto et al. (2012) already noted that phylogenetic distance can be a proxy of niche overlap in tropical tree communities. Here, Schmitt et al. further demonstrate that evolutionary heritage is significantly related to individual growth variation, and plead for better acknowledging this role in future studies. While the role of fitness differences in tropical tree community dynamics remained to be assessed, the present study provides new evidence that individual growth does vary depending on evolutionary relationships, which can reflect the roles of selection and adaptation on growth strategies. Therefore, investigating both the influence of functional traits and phylogenetic relationships on individual performance remains a promising avenue of research, for functional and community ecology in general. REFERENCES Baraloto, Christopher, Olivier J. Hardy, C. E. Timothy Paine, Kyle G. Dexter, Corinne Cruaud, Luke T. Dunning, Mailyn-Adriana Gonzalez, et al. 2012. « Using functional traits and phylogenetic trees to examine the assembly of tropical tree communities ». Journal of Ecology, 100: 690‑701. | High intraspecific growth variability despite strong evolutionary heritage in a neotropical forest | Sylvain Schmitt, Bruno Hérault, Géraldine Derroire | <p style="text-align: justify;">Individual tree growth is a key determinant of species performance and a driver of forest dynamics and composition. Previous studies on tree growth unravelled the variation in species growth as a function of demogra... |  | Community ecology, Demography, Population ecology | François Munoz | Jelena Pantel, David Murray-Stoker | 2022-08-01 14:29:04 | View |
30 May 2024
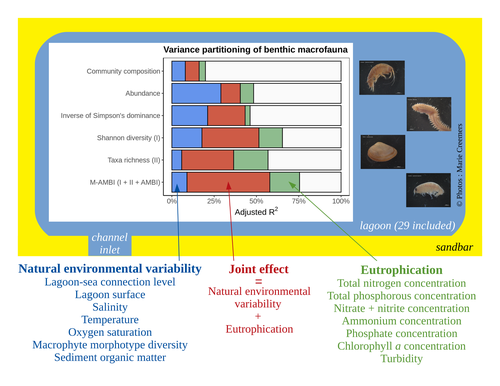
Disentangling the effects of eutrophication and natural variability on macrobenthic communities across French coastal lagoonsAuriane G. Jones, Gauthier Schaal, Aurélien Boyé, Marie Creemers, Valérie Derolez, Nicolas Desroy, Annie Fiandrino, Théophile L. Mouton, Monique Simier, Niamh Smith, Vincent Ouisse https://doi.org/10.1101/2022.08.18.504439Untangling Eutrophication Effects on Coastal Lagoon EcosystemsRecommended by Nathalie Niquil based on reviews by Kaylee P. Smit, Matthew J. Pruden and Kendyl Wright based on reviews by Kaylee P. Smit, Matthew J. Pruden and Kendyl Wright
Disentangling the effects on ecosystem structure and functioning of natural and human-induced impacts in transitional waters is a great challenge in coast ecology. This is due to the observation that the ecosystems of transitional waters are naturally dynamic systems with characteristics of stressed systems. For example, the benthic communities present low species richness and high abundance of species with a high tolerance to variations, e.g., salinity. This general observation is known as the paradigm of the “Transitional Waters Quality Paradox” (Zaldívar et al., 2008) derived from the previously described “Estuarine Quality Paradox” (Elliott and Quintino, 2007). In Jones et al. (2024) “Disentangling the effects of eutrophication and natural variability on macrobenthic communities across French coastal lagoons”, a great diversity of lagoons is analyzed to disentangle the effects of eutrophication from those of natural environmental variability on benthic macroinvertebrates and understanding the links between environmental variables affecting benthic macroinvertebrates. These authors use a very elegant set of numerical approaches, including correlograms, linear models and variance partitioning. They apply this suite to a dataset of macrobenthic invertebrate abundances and environmental variables from 29 Mediterranean coastal lagoons in France. Through this suite of analyses, they demonstrate the strong complexity of the mechanisms interplaying in a situation of eutrophication on lagoon macrobenthos. The mechanisms involved are direct, like toxicity, or indirect, for example, through modifications of the sediment's biogeochemistry. Such a result on the different interactions involved is very important in the context of the search for indicators to define ecosystem status. Improving the definition of metrics is essential in environmental management decisions. References Elliott, M. and Quintino, V. (2007) The estuarine quality paradox, environmental homeostasis and the difficulty of detecting anthropogenic stress in naturally stressed areas. Marine Pollution Bulletin 54, 640–645. https://doi.org/10.1016/j.marpolbul.2007.02.003 Zaldívar, J. (2008). Eutrophication in transitional waters: an overview. https://doi.org/10.1285/I18252273V2N1P1 | Disentangling the effects of eutrophication and natural variability on macrobenthic communities across French coastal lagoons | Auriane G. Jones, Gauthier Schaal, Aurélien Boyé, Marie Creemers, Valérie Derolez, Nicolas Desroy, Annie Fiandrino, Théophile L. Mouton, Monique Simier, Niamh Smith, Vincent Ouisse | <p style="text-align: justify;">Coastal lagoons are transitional ecosystems that host a unique diversity of species and support many ecosystem services. Owing to their position at the interface between land and sea, they are also subject to increa... |  | Biodiversity, Community ecology, Ecosystem functioning, Marine ecology | Nathalie Niquil | Matthew J. Pruden | 2023-09-08 11:26:01 | View |
09 Dec 2019

Niche complementarity among pollinators increases community-level plant reproductive successAinhoa Magrach, Francisco P. Molina, Ignasi Bartomeus https://doi.org/10.1101/629931Improving our knowledge of species interaction networksRecommended by Cédric Gaucherel based on reviews by Michael Lattorff, Nicolas Deguines and 3 anonymous reviewers based on reviews by Michael Lattorff, Nicolas Deguines and 3 anonymous reviewers
Ecosystems shelter a huge number of species, continuously interacting. Each species interact in various ways, with trophic interactions, but also non-trophic interactions, not mentioning the abiotic and anthropogenic interactions. In particular, pollination, competition, facilitation, parasitism and many other interaction types are simultaneously present at the same place in terrestrial ecosystems [1-2]. For this reason, we need today to improve our understanding of such complex interaction networks to later anticipate their responses. This program is a huge challenge facing ecologists and they today join their forces among experimentalists, theoreticians and modelers. While some of us struggle in theoretical and modeling dimensions [3-4], some others perform brilliant works to observe and/or experiment on the same ecological objects [5-6]. References [1] Campbell, C., Yang, S., Albert, R., and Shea, K. (2011). A network model for plant–pollinator community assembly. Proceedings of the National Academy of Sciences, 108(1), 197-202. doi: 10.1073/pnas.1008204108 | Niche complementarity among pollinators increases community-level plant reproductive success | Ainhoa Magrach, Francisco P. Molina, Ignasi Bartomeus | <p>Declines in pollinator diversity and abundance have been reported across different regions, with implications for the reproductive success of plant species. However, research has focused primarily on pairwise plant-pollinator interactions, larg... |  | Ecosystem functioning, Interaction networks, Pollination, Terrestrial ecology | Cédric Gaucherel | Nicolas Deguines | 2019-05-07 17:03:23 | View |
24 May 2023
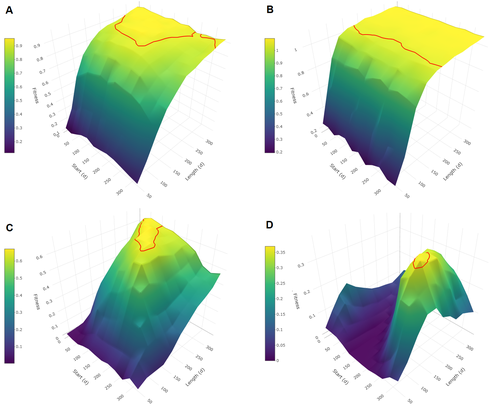
Evolutionary determinants of reproductive seasonality: a theoretical approachLugdiwine Burtschell, Jules Dezeure, Elise Huchard, Bernard Godelle https://doi.org/10.1101/2022.08.22.504761When does seasonal reproduction evolve?Recommended by Tim Coulson based on reviews by Francois-Xavier Dechaume-Moncharmont, Nigel Yoccoz and 1 anonymous reviewer based on reviews by Francois-Xavier Dechaume-Moncharmont, Nigel Yoccoz and 1 anonymous reviewer
Have you ever wondered why some species breed seasonally while others do not? You might think it is all down to lattitude and the harshness of winters but it turns out it is quite a bit more complicated than that. A consequence of this is that climate change may result in the evolution of the degree of seasonal reproduction, with some species perhaps becoming less seasonal and others more so even in the same habitat. Burtschell et al. (2023) investigated how various factors influence seasonal breeding by building an individual-based model of a baboon population from which they calculated the degree of seasonality for the fittest reproductive strategy. They then altered key aspects of their model to examine how these changes impacted the degree of seasonality in the reproductive strategy. What they found is fascinating. The degree of seasonality in reproductive strategy is expected to increase with increased seasonality in the environment, decreased food availability, increased energy expenditure, and how predictable resource availability is. Interestingly, neither female cycle length nor extrinsic infant mortality influenced the degree of seasonality in reproduction. What this means in reality for seasonal species is more challenging to understand. Some environments appear to be becoming more seasonal yet less predictable, and some species appear to be altering their daily energy budgets in response to changing climate in quite complex ways. As with pretty much everything in biology, Burtschell et al.'s work reveals much nuance and complexity, and that predicting how species might alter their reproductive timing is fraught with challenges. The paper is very well written. With a simpler model it may have proven possible to achieve analytical solutions, but this is a very minor gripe. The reviewers were positive about the paper, and I have little doubt it will be well-cited. REFERENCES Burtschell L, Dezeure J, Huchard E, Godelle B (2023) Evolutionary determinants of reproductive seasonality: a theoretical approach. bioRxiv, 2022.08.22.504761, ver. 2 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2022.08.22.504761 | Evolutionary determinants of reproductive seasonality: a theoretical approach | Lugdiwine Burtschell, Jules Dezeure, Elise Huchard, Bernard Godelle | <p style="text-align: justify;">Reproductive seasonality is a major adaptation to seasonal cycles and varies substantially among organisms. This variation, which was long thought to reflect a simple latitudinal gradient, remains poorly understood ... |  | Evolutionary ecology, Life history, Theoretical ecology | Tim Coulson | Nigel Yoccoz | 2022-08-23 21:37:28 | View |
10 Oct 2018

Detecting within-host interactions using genotype combination prevalence dataSamuel Alizon, Carmen Lía Murall, Emma Saulnier, Mircea T Sofonea https://doi.org/10.1101/256586Combining epidemiological models with statistical inference can detect parasite interactionsRecommended by Dustin Brisson based on reviews by Samuel Díaz Muñoz, Erick Gagne and 1 anonymous reviewerThere are several important topics in the study of infectious diseases that have not been well explored due to technical difficulties. One such topic is pursued by Alizon et al. in “Modelling coinfections to detect within-host interactions from genotype combination prevalences” [1]. Both theory and several important examples have demonstrated that interactions among co-infecting strains can have outsized impacts on disease outcomes, transmission dynamics, and epidemiology. Unfortunately, empirical data on pathogen interactions and their outcomes is often correlational making results difficult to decipher. References [1] Alizon, S., Murall, C.L., Saulnier, E., & Sofonea, M.T. (2018). Detecting within-host interactions using genotype combination prevalence data. bioRxiv, 256586, ver. 3 peer-reviewed and recommended by PCI Ecology. doi: 10.1101/256586 | Detecting within-host interactions using genotype combination prevalence data | Samuel Alizon, Carmen Lía Murall, Emma Saulnier, Mircea T Sofonea | <p>Parasite genetic diversity can provide information on disease transmission dynamics but most methods ignore the exact combinations of genotypes in infections. We introduce and validate a new method that combines explicit epidemiological modelli... |  | Eco-immunology & Immunity, Epidemiology, Host-parasite interactions, Statistical ecology | Dustin Brisson | Samuel Díaz Muñoz, Erick Gagne | 2018-02-01 09:23:26 | View |
24 May 2024
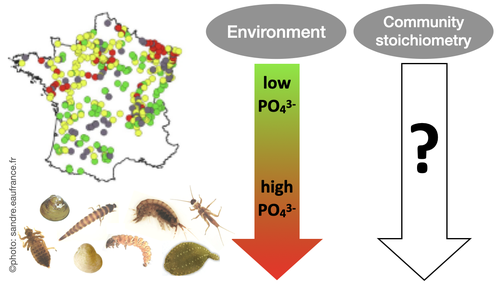
Effects of water nutrient concentrations on stream macroinvertebrate community stoichiometry: a large-scale studyMiriam Beck, Elise Billoir, Philippe Usseglio-Polatera, Albin Meyer, Edwige Gautreau, Michael Danger https://doi.org/10.1101/2024.02.01.574823The influence of water phosphorus and nitrogen loads on stream macroinvertebrate community stoichiometryRecommended by Huihuang Chen based on reviews by Thomas Guillemaud, Jun Zuo and 1 anonymous reviewer based on reviews by Thomas Guillemaud, Jun Zuo and 1 anonymous reviewer
The manuscript by Beck et al. (2024) investigates the effects of water phosphorus and nitrogen loads on stream macroinvertebrate community stoichiometry across France. Utilizing data from over 1300 standardized sampling events, this research finds that community stoichiometry is significantly influenced by water phosphorus concentration, with the strongest effects at low nitrogen levels. The results demonstrate that the assumptions of Ecological Stoichiometry Theory apply at the community level for at least two dominant taxa and across a broad spatial scale, with probable implications for nutrient cycling and ecosystem functionality. This manuscript contributes to ecological theory, particularly by extending Ecological Stoichiometry Theory to include community-level interactions, clarifying the impact of nutrient concentrations on community structure and function, and informing nutrient management and conservation strategies. In summary, this study not only addresses a gap in community-level stoichiometric research but also delivers crucial empirical support for advancing ecological science and promoting environmental stewardship. References Beck M, Billoir E, Usseglio-Polatera P, Meyer A, Gautreau E and Danger M (2024) Effects of water nutrient concentrations on stream macroinvertebrate community stoichiometry: a large-scale study. bioRxiv, 2024.02.01.574823, ver. 2 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2024.02.01.574823 | Effects of water nutrient concentrations on stream macroinvertebrate community stoichiometry: a large-scale study | Miriam Beck, Elise Billoir, Philippe Usseglio-Polatera, Albin Meyer, Edwige Gautreau, Michael Danger | <p>Basal resources generally mirror environmental nutrient concentrations in the elemental composition of their tissue, meaning that nutrient alterations can directly reach consumer level. An increased nutrient content (e.g. phosphorus) in primary... |  | Community ecology, Ecological stoichiometry | Huihuang Chen | Thomas Guillemaud, Jun Zuo, Anonymous | 2024-02-02 10:14:01 | View |
03 Jan 2024
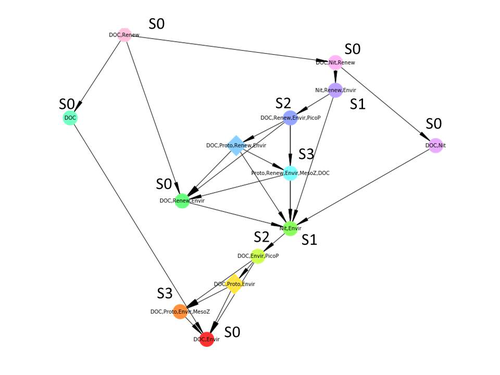
Diagnosis of planktonic trophic network dynamics with sharp qualitative changesCedric Gaucherel, Stolian Fayolle, Raphael Savelli, Olivier Philippine, Franck Pommereau, Christine Dupuy https://doi.org/10.1101/2023.06.29.547055A new approach to describe qualitative changes of complex trophic networksRecommended by Francis Raoul based on reviews by Tim Coulson and 1 anonymous reviewerModelling the temporal dynamics of trophic networks has been a key challenge for community ecologists for decades, especially when anthropogenic and natural forces drive changes in species composition, abundance, and interactions over time. So far, most modelling methods fail to incorporate the inherent complexity of such systems, and its variability, to adequately describe and predict temporal changes in the topology of trophic networks. Taking benefit from theoretical computer science advances, Gaucherel and colleagues (2024) propose a new methodological framework to tackle this challenge based on discrete-event Petri net methodology. To introduce the concept to naïve readers the authors provide a useful example using a simplistic predator-prey model. The core biological system of the article is a freshwater trophic network of western France in the Charente-Maritime marshes of the French Atlantic coast. A directed graph describing this system was constructed to incorporate different functional groups (phytoplankton, zooplankton, resources, microbes, and abiotic components of the environment) and their interactions. Rules and constraints were then defined to, respectively, represent physiochemical, biological, or ecological processes linking network components, and prevent the model from simulating unrealistic trajectories. Then the full range of possible trajectories of this mechanistic and qualitative model was computed. The model performed well enough to successfully predict a theoretical trajectory plus two trajectories of the trophic network observed in the field at two different stations, therefore validating the new methodology introduced here. The authors conclude their paper by presenting the power and drawbacks of such a new approach to qualitatively model trophic networks dynamics. Reference Cedric Gaucherel, Stolian Fayolle, Raphael Savelli, Olivier Philippine, Franck Pommereau, Christine Dupuy (2024) Diagnosis of planktonic trophic network dynamics with sharp qualitative changes. bioRxiv 2023.06.29.547055, ver. 2 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2023.06.29.547055 | Diagnosis of planktonic trophic network dynamics with sharp qualitative changes | Cedric Gaucherel, Stolian Fayolle, Raphael Savelli, Olivier Philippine, Franck Pommereau, Christine Dupuy | <p>Trophic interaction networks are notoriously difficult to understand and to diagnose (i.e., to identify contrasted network functioning regimes). Such ecological networks have many direct and indirect connections between species, and these conne... |  | Community ecology, Ecosystem functioning, Food webs, Freshwater ecology, Interaction networks, Microbial ecology & microbiology | Francis Raoul | Tim Coulson | 2023-07-03 10:42:34 | View |
23 Oct 2023
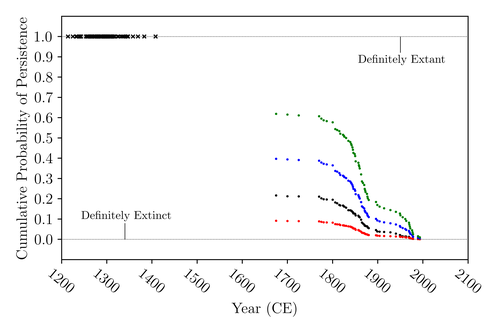
The Moa the Merrier: Resolving When the Dinornithiformes Went ExtinctFloe Foxon https://doi.org/10.1101/2023.08.07.552261Are Moas ancient Lazarus species?Recommended by Werner Ulrich based on reviews by Tim Coulson and Richard Holdaway based on reviews by Tim Coulson and Richard Holdaway
Ancient human colonisation often had catastrophic consequences for native fauna. The North American Megafauna went extinct shortly after humans entered the scene and Madagascar suffered twice, before 1500 CE and around 1700 CE after the Malayan and European colonisation. Maoris colonised New Zealand by about 1300 and a century later the giant Moa birds (Dinornithiformes) sharply declined. But did they went extinct or are they an ancient example of Lazarus species, species thought to be extinct but still alive? Scattered anecdotes of late sightings of living Moas even up to the 20th century seem to suggest the latter. The quest for later survival has also a criminal aspect. Who did it, the Maoris or the white colonisers in the late 18th century? The present work by Floe Foxon (2023) tries to settle this question. It uses a survival modelling approach and an assessment of the reliability of nearly 100 alleged sightings. The model favours the so-called overkill hypothesis, that Moas probably went extinct in the 15th century shortly after Maori colonisation. A small but still remarkable probability remained for survival up to 1770. Later sightings turned out to be highly unreliable. The paper is important as it does not rely on subjective discussions of late sightings but on a probabilistic modelling approach with sensitivity testing prior applied to marsupials. As common in probabilistic approaches, the study does not finally settle the case. A probability of as much as 20% remained for late survival after 1450 CE. This is not improbable as New Zealand was sufficiently unexplored in those days to harbour a few refuges for late survivors. However, in this respect, it is a bit unfortunate that at the end of the discussion, the paper cites Heuvelmans, the founder of cryptozoology, and it mentions the ivory-billed woodpecker, which has recently been redetected. No Moa remains were found after 1450. References Foxon F (2023) The Moa the Merrier: Resolving When the Dinornithiformes Went Extinct. bioRxiv, 2023.08.07.552261, ver. 2 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2023.08.07.552261 | The Moa the Merrier: Resolving When the Dinornithiformes Went Extinct | Floe Foxon | <p style="text-align: justify;">The Moa (Aves: Dinornithiformes) are an extinct group of the ratite clade from New Zealand. The overkill hypothesis asserts that the first New Zealand settlers hunted the Moa to extinction by 1450 CE, whereas the st... |  | Conservation biology, Human impact, Statistical ecology, Zoology | Werner Ulrich | Tim Coulson, Richard Holdaway | 2023-08-08 17:14:30 | View |
20 Sep 2018
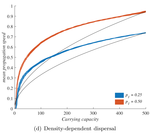
When higher carrying capacities lead to faster propagationMarjorie Haond, Thibaut Morel-Journel, Eric Lombaert, Elodie Vercken, Ludovic Mailleret & Lionel Roques https://doi.org/10.1101/307322When the dispersal of the many outruns the dispersal of the fewRecommended by Matthieu Barbier based on reviews by Yuval Zelnik and 1 anonymous reviewer based on reviews by Yuval Zelnik and 1 anonymous reviewer
Are biological invasions driven by a few pioneers, running ahead of their conspecifics? Or are these pioneers constantly being caught up by, and folded into, the larger flux of propagules from the established populations behind them? References [1] Levins, R., & Culver, D. (1971). Regional Coexistence of Species and Competition between Rare Species. Proceedings of the National Academy of Sciences, 68(6), 1246–1248. doi: 10.1073/pnas.68.6.1246 | When higher carrying capacities lead to faster propagation | Marjorie Haond, Thibaut Morel-Journel, Eric Lombaert, Elodie Vercken, Ludovic Mailleret & Lionel Roques | <p>This preprint has been reviewed and recommended by Peer Community In Ecology (https://dx.doi.org/10.24072/pci.ecology.100004). Finding general patterns in the expansion of natural populations is a major challenge in ecology and invasion biology... |  | Biological invasions, Colonization, Dispersal & Migration, Experimental ecology, Population ecology, Spatial ecology, Metacommunities & Metapopulations, Theoretical ecology | Matthieu Barbier | Yuval Zelnik | 2018-04-25 10:18:48 | View |
MANAGING BOARD
Julia Astegiano
Tim Coulson
Anna Eklof
Dominique Gravel
François Massol
Ben Phillips
Cyrille Violle










