Direct submissions to PCI Ecology from bioRxiv.org are possible using the B2J service
Latest recommendations

| Id | Title * | Authors * | Abstract * | Picture * | Thematic fields * | Recommender▲ | Reviewers | Submission date | |
|---|---|---|---|---|---|---|---|---|---|
09 Nov 2023
Mark loss can strongly bias estimates of demographic rates in multi-state models: a case study with simulated and empirical datasetsFrédéric Touzalin, Eric J. Petit, Emmanuelle Cam, Claire Stagier, Emma C. Teeling, Sébastien J. Puechmaille https://doi.org/10.1101/2022.03.25.485763Marks lost in action, biased estimationsRecommended by Sylvain Billiard based on reviews by Olivier Gimenez, Devin Johnson and 1 anonymous reviewer based on reviews by Olivier Gimenez, Devin Johnson and 1 anonymous reviewer
Capture-Mark-Recapture (CMR) data are commonly used to estimate ecological variables such as abundance, survival probability, or transition rates from one state to another (e.g. from juvenile to adult, or migration from one site to another). Many studies have shown how estimations can be affected by neglecting one aspect of the population under study (e.g. the heterogeneity in survival between individuals) or one limit of the methodology itself (e.g. the fact that observers might not detect an individual although it is still alive). Strikingly, very few studies have yet assessed the robustness of one fundamental assumption of all CMR-based inferences: marks are supposed definitive and immutable. If they are not, how are estimations affected? Addressing this issue is the main goal of the paper by Touzalin et al. (2023), and they did a very nice work. But, because the answer is not that simple, it also calls for further investigations. When and why would mark loss bias estimation? In at least two situations. First, when estimating survival rates: if an individual loses its mark, it will be considered as dead, hence death rates will be overestimated. Second, more subtly, when estimating transition rates: if one individual loses its mark at the specific moment where its state changes, then a transition will be missed in data. The history of the marked individual would then be split into two independent CMR sequences as if there were two different individuals, including one which died. Touzalin et al. (2023) thoroughly studied these two situations by estimating ecological parameters on 1) well-thought simulated datasets, that cover a large range of possible situations inspired from a nice compilation of hundreds of estimations from fish and bats studies, and 2) on their own bats dataset, for which they had various sources of information about mark losses, i.e. different mark types on the same individuals, including mark based on genotypes, and marks found on the soil in the place where bats lived. Their main findings from the simulated datasets are that there is a general trend for underestimation of survival and transition rates if mark loss is not accounting for in the model, as it would be intuitively expected. However, they also showed from the bats dataset that biases do not show any obvious general trend, suggesting complex interactions between different ecological processes and/or with the estimation procedure itself. The results by Touzalin et al. (2023) strongly suggest that mark loss should systematically be included in models estimating parameters from CMR data. In addition to adapt the inferential models, the authors also recommend considering either a double marking, or even a single but ‘permanent’ mark such as one based on the genotypes. However, the potential gain of a double marking or of the use of genotypes is still to be evaluated both in theory and practice, and it seems to be not that obvious at first sight. First because double marking can be costly for experimenters but also for the marked animals, especially as several studies showed that marks can significantly affect survival or recapture rates. Second because multiple sources of errors can affect genotyping, which would result in wrong individual assignations especially in populations with low genetic diversity or high inbreeding, or no individual assignation at all, which would increase the occurrence of missing data in CMR datasets. Touzalin et al. (2023) supposed in their paper that there were no genotyping errors, but one can doubt it to be true in most situations. They have now important and interesting other issues to address. References Frédéric Touzalin, Eric J. Petit, Emmanuelle Cam, Claire Stagier, Emma C. Teeling, Sébastien J. Puechmaille (2023) Mark loss can strongly bias demographic rates in multi-state models: a case study with simulated and empirical datasets. BioRxiv, ver. 3 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2022.03.25.485763 | Mark loss can strongly bias estimates of demographic rates in multi-state models: a case study with simulated and empirical datasets | Frédéric Touzalin, Eric J. Petit, Emmanuelle Cam, Claire Stagier, Emma C. Teeling, Sébastien J. Puechmaille | <p style="text-align: justify;">1. The development of methods for individual identification in wild species and the refinement of Capture-Mark-Recapture (CMR) models over the past few decades have greatly improved the assessment of population demo... | Conservation biology, Demography | Sylvain Billiard | 2022-04-12 18:49:34 | View | ||
05 Jun 2024
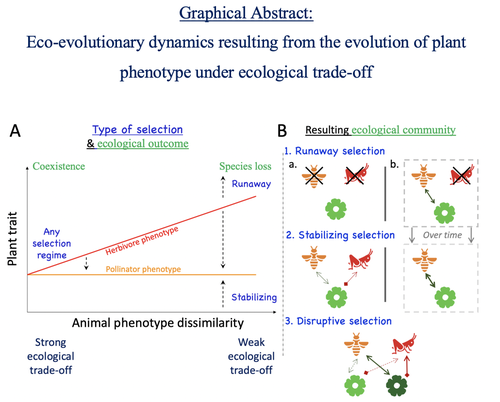
Attracting pollinators vs escaping herbivores: eco-evolutionary dynamics of plants confronted with an ecological trade-offYoussef Yacine, Nicolas Loeuille https://doi.org/10.1101/2021.12.02.470900Plant-herbivore-pollinator ménage-à-trois: tell me how well they match, and I'll tell you if it's made to lastRecommended by Sylvain Billiard based on reviews by Marcos Mendez and Yaroslav Ispolatov based on reviews by Marcos Mendez and Yaroslav Ispolatov
How would a plant trait evolve if it is involved in interacting with both a pollinator and an herbivore species? The answer by Yacine and Loeuille is straightforward: it is not trivial, but it can explain many situations found in natural populations. Yacine and Loeuille applied the well-known Adaptive Dynamics framework to a system with three interacting protagonists: a herbivore, a pollinator, and a plant. The evolution of a plant trait is followed under the assumption that it regulates the frequency of interaction with the two other species. As one can imagine, that is where problems begin: interacting more with pollinators seems good, but what if at the same time it implies interacting more with herbivores? And that's not a silly idea, as there are many cases where herbivores and pollinators share the same cues to detect plants, such as colors or chemical compounds. They found that depending on the trade-off between the two types of interactions and their density-dependent effects on plant fitness, the possible joint ecological and evolutionary outcomes are numerous. When herbivory prevails, evolution can make the ménage-à-trois ecologically unstable, as one or even two species can go extinct, leaving the plant alone. Evolution can also make the coexistence of the three species more stable when pollination services prevail, or lead to the appearance of a second plant species through branching diversification of the plant trait when herbivory and pollination are balanced. Yacine and Loeuille did not only limit themselves to saying "it is possible," but they also did much work evaluating when each evolutionary outcome would occur. They numerically explored in great detail the adaptive landscape of the plant trait for a large range of parameter values. They showed that the global picture is overall robust to parameter variations, strengthening the plausibility that the evolution of a trait involved in antagonistic interactions can explain many of the correlations between plant and animal traits or phylogenies found in nature. Are we really there yet? Of course not, as some assumptions of the model certainly limit its scope. Are there really cases where plants' traits evolve much faster than herbivores' and pollinators' traits? Certainly not, but the model is so general that it can apply to any analogous system where one species is caught between a mutualistic and a predator species, including potential species that evolve much faster than the two others. And even though this limitation might cast doubt on the generality of the model's predictions, studying a system where a species' trait and a preference trait coevolve is possible, as other models have already been studied (see Fritsch et al. 2021 for a review in the case of evolution in food webs). We can bet this is the next step taken by Yacine and Loeuille in a similar framework with the same fundamental model, promising fascinating results, especially regarding the evolution of complex communities when species can accumulate after evolutionary branchings. Relaxing another assumption seems more challenging as it would certainly need to change the model itself: interacting species generally do not play fixed roles, as being mutualistic or antagonistic might generally be density-dependent (Holland and DeAngelis 2010). How would the exchange of resources between three interacting species evolve? It is an open question. References Fritsch, C., Billiard, S., & Champagnat, N. (2021). Identifying conversion efficiency as a key mechanism underlying food webs adaptive evolution: a step forward, or backward? Oikos, 130(6), 904-930. Yacine, Y., & Loeuille, N. (2024) Attracting pollinators vs escaping herbivores: eco-evolutionary dynamics of plants confronted with an ecological trade-off. bioRxiv 2021.12.02.470900; doi: https://doi.org/10.1101/2021.12.02.470900 | Attracting pollinators vs escaping herbivores: eco-evolutionary dynamics of plants confronted with an ecological trade-off | Youssef Yacine, Nicolas Loeuille | <p style="text-align: justify;">Many plant traits are subject to an ecological trade-off between attracting pollinators and escaping herbivores. The interplay of both plant-animal interaction types determines their evolution. As most studies focus... |  | Eco-evolutionary dynamics, Herbivory, Pollination, Theoretical ecology | Sylvain Billiard | 2023-03-21 14:23:12 | View | |
16 Sep 2019
Blood, sweat and tears: a review of non-invasive DNA samplingMarie-Caroline Lefort, Robert H Cruickshank, Kris Descovich, Nigel J Adams, Arijana Barun, Arsalan Emami-Khoyi, Johnaton Ridden, Victoria R Smith, Rowan Sprague, Benjamin Waterhouse, Stephane Boyer https://doi.org/10.1101/385120Words matter: extensive misapplication of "non-invasive" in describing DNA sampling methods, and proposed clarifying termsRecommended by Thomas Wilson Sappington based on reviews by 2 anonymous reviewersThe ability to successfully sequence trace quantities of environmental DNA (eDNA) has provided unprecedented opportunities to use genetic analyses to elucidate animal ecology, behavior, and population structure without affecting the behavior, fitness, or welfare of the animal sampled. Hair associated with an animal track in the snow, the shed exoskeleton of an insect, or a swab of animal scat are all examples of non-invasive methods to collect eDNA. Despite the seemingly uncomplicated definition of "non-invasive" as proposed by Taberlet et al. [1], Lefort et al. [2] highlight that its appropriate application to sampling methods in practice is not so straightforward. For example, collecting scat left behind on the forest floor by a mammal could be invasive if feces is used by that species to mark territorial boundaries. Other collection strategies such as baited DNA traps to collect hair, capturing and handling an individual to swab or stimulate emission of a body fluid, or removal of a presumed non essential body part like a feather, fish scale, or even a leg from an insect are often described as "non-invasive" sampling methods. However, such methods cannot be considered truly non-invasive. At a minimum, attracting or capturing and handling an animal to obtain a DNA sample interrupts its normal behavioral routine, but additionally can cause both acute and long-lasting physiological and behavioral stress responses and other effects. Even invertebrates exhibit long-term hypersensitization after an injury, which manifests as heightened vigilance and enhanced escape responses [3-5]. References [1] Taberlet P., Waits L. P. and Luikart G. 1999. Noninvasive genetic sampling: look before you leap. Trends Ecol. Evol. 14: 323-327. doi: 10.1016/S0169-5347(99)01637-7 | Blood, sweat and tears: a review of non-invasive DNA sampling | Marie-Caroline Lefort, Robert H Cruickshank, Kris Descovich, Nigel J Adams, Arijana Barun, Arsalan Emami-Khoyi, Johnaton Ridden, Victoria R Smith, Rowan Sprague, Benjamin Waterhouse, Stephane Boyer | <p>The use of DNA data is ubiquitous across animal sciences. DNA may be obtained from an organism for a myriad of reasons including identification and distinction between cryptic species, sex identification, comparisons of different morphocryptic ... |  | Behaviour & Ethology, Conservation biology, Molecular ecology, Zoology | Thomas Wilson Sappington | 2018-11-30 13:33:31 | View | |
24 May 2022
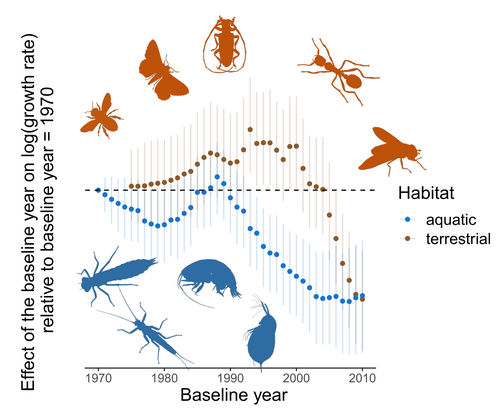
Controversy over the decline of arthropods: a matter of temporal baseline?François Duchenne, Emmanuelle Porcher, Jean-Baptiste Mihoub, Grégoire Loïs, Colin Fontaine https://doi.org/10.1101/2022.02.09.479422Don't jump to conclusions on arthropod abundance dynamics without appropriate dataRecommended by Tim Coulson based on reviews by Gabor L Lovei and 1 anonymous reviewer based on reviews by Gabor L Lovei and 1 anonymous reviewer
Humans are dramatically modifying many aspects of our planet via increasing concentrations of carbon dioxide in the atmosphere, patterns of land-use change, and unsustainable exploitation of the planet’s resources. These changes impact the abundance of species of wild organisms, with winners and losers. Identifying how different species and groups of species are influenced by anthropogenic activity in different biomes, continents, and habitats, has become a pressing scientific question with many publications reporting analyses of disparate data on species population sizes. Many conclusions are based on the linear analysis of rather short time series of organismal abundances. Duchenne F, Porcher E, Mihoub J-B, Loïs G, Fontaine C (2022) Controversy over the decline of arthropods: a matter of temporal baseline? bioRxiv, 2022.02.09.479422, ver. 3 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2022.02.09.479422 | Controversy over the decline of arthropods: a matter of temporal baseline? | François Duchenne, Emmanuelle Porcher, Jean-Baptiste Mihoub, Grégoire Loïs, Colin Fontaine | <p style="text-align: justify;">Recently, a number of studies have reported somewhat contradictory patterns of temporal trends in arthropod abundance, from decline to increase. Arthropods often exhibit non-monotonous variation in abundance over ti... |  | Conservation biology | Tim Coulson | 2022-02-11 15:44:44 | View | |
03 Jun 2022
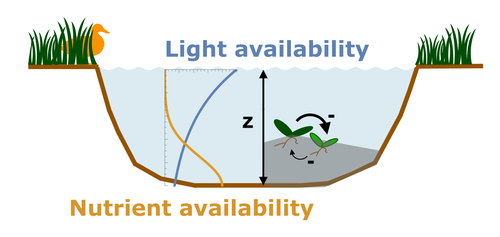
Evolutionary emergence of alternative stable states in shallow lakesAlice Ardichvili, Nicolas Loeuille, Vasilis Dakos https://doi.org/10.1101/2022.02.23.481597How to evolve an alternative stable stateRecommended by Tim Coulson based on reviews by Jean-François Arnoldi and 1 anonymous reviewer based on reviews by Jean-François Arnoldi and 1 anonymous reviewer
Alternative stable states describe ecosystems that can persist in more than one configuration. An ecosystem can shift between stable states following some form of perturbation. There has been much work on predicting when ecosystems will shift between stable states, but less work on why some ecosystems are able to exist in alternative stable states in the first place. The paper by Ardichvili, Loeuille, and Dakos (2022) addresses this question using a simple model of a shallow lake. Their model is based on a trade-off between access to light and nutrient availability in the water column, two essential resources for the macrophytes they model. They then identify conditions when the ancestral macrophyte will diversify resulting in macrophyte species living at new depths within the lake. The authors find a range of conditions where alternative stable states can evolve, but the range is narrow. Nonetheless, their model suggests that for alternative stable states to exist, one requirement is for there to be asymmetric competition between competing species, with one species being a better competitor on one limiting resource, with the other being a better competitor on a second limiting resource. These results are interesting and add to growing literature on how asymmetric competition can aid species coexistence. Asymmetric competition may be widespread in nature, with closely related species often being superior competitors on different resources. Incorporating asymmetric competition, and its evolution, into models does complicate theoretical investigations, but Ardichvili, Loeuille, and Dakos’ paper elegantly shows how substantial progress can be made with a model that is still (relatively) simple. References Ardichvili A, Loeuille N, Dakos V (2022) Evolutionary emergence of alternative stable states in shallow lakes. bioRxiv, 2022.02.23.481597, ver. 3 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2022.02.23.481597 | Evolutionary emergence of alternative stable states in shallow lakes | Alice Ardichvili, Nicolas Loeuille, Vasilis Dakos | <p style="text-align: justify;">Ecosystems under stress may respond abruptly and irreversibly through tipping points. Although much is explored on the mechanisms that affect tipping points and alternative stable states, little is known on how ecos... |  | Community ecology, Competition, Eco-evolutionary dynamics, Theoretical ecology | Tim Coulson | 2022-03-01 10:54:05 | View | |
24 May 2023
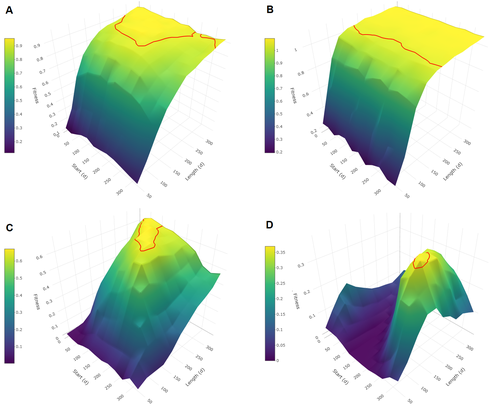
Evolutionary determinants of reproductive seasonality: a theoretical approachLugdiwine Burtschell, Jules Dezeure, Elise Huchard, Bernard Godelle https://doi.org/10.1101/2022.08.22.504761When does seasonal reproduction evolve?Recommended by Tim Coulson based on reviews by Francois-Xavier Dechaume-Moncharmont, Nigel Yoccoz and 1 anonymous reviewer based on reviews by Francois-Xavier Dechaume-Moncharmont, Nigel Yoccoz and 1 anonymous reviewer
Have you ever wondered why some species breed seasonally while others do not? You might think it is all down to lattitude and the harshness of winters but it turns out it is quite a bit more complicated than that. A consequence of this is that climate change may result in the evolution of the degree of seasonal reproduction, with some species perhaps becoming less seasonal and others more so even in the same habitat. Burtschell et al. (2023) investigated how various factors influence seasonal breeding by building an individual-based model of a baboon population from which they calculated the degree of seasonality for the fittest reproductive strategy. They then altered key aspects of their model to examine how these changes impacted the degree of seasonality in the reproductive strategy. What they found is fascinating. The degree of seasonality in reproductive strategy is expected to increase with increased seasonality in the environment, decreased food availability, increased energy expenditure, and how predictable resource availability is. Interestingly, neither female cycle length nor extrinsic infant mortality influenced the degree of seasonality in reproduction. What this means in reality for seasonal species is more challenging to understand. Some environments appear to be becoming more seasonal yet less predictable, and some species appear to be altering their daily energy budgets in response to changing climate in quite complex ways. As with pretty much everything in biology, Burtschell et al.'s work reveals much nuance and complexity, and that predicting how species might alter their reproductive timing is fraught with challenges. The paper is very well written. With a simpler model it may have proven possible to achieve analytical solutions, but this is a very minor gripe. The reviewers were positive about the paper, and I have little doubt it will be well-cited. REFERENCES Burtschell L, Dezeure J, Huchard E, Godelle B (2023) Evolutionary determinants of reproductive seasonality: a theoretical approach. bioRxiv, 2022.08.22.504761, ver. 2 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2022.08.22.504761 | Evolutionary determinants of reproductive seasonality: a theoretical approach | Lugdiwine Burtschell, Jules Dezeure, Elise Huchard, Bernard Godelle | <p style="text-align: justify;">Reproductive seasonality is a major adaptation to seasonal cycles and varies substantially among organisms. This variation, which was long thought to reflect a simple latitudinal gradient, remains poorly understood ... |  | Evolutionary ecology, Life history, Theoretical ecology | Tim Coulson | Nigel Yoccoz | 2022-08-23 21:37:28 | View |
31 Aug 2023
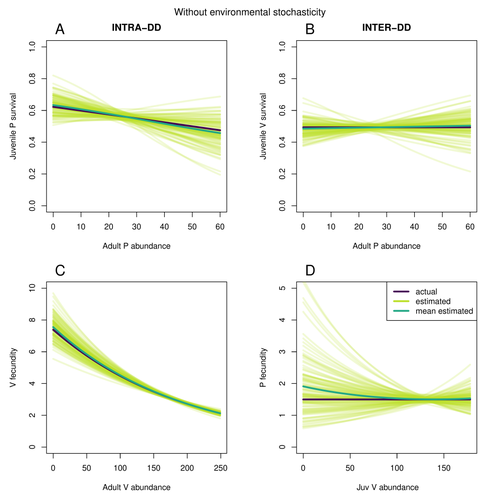
Assessing species interactions using integrated predator-prey modelsMatthieu Paquet, Frederic Barraquand https://doi.org/10.32942/X2RC7WAddressing the daunting challenge of estimating species interactions from count dataRecommended by Tim Coulson and David Alonso and David Alonso based on reviews by 2 anonymous reviewers based on reviews by 2 anonymous reviewers
Trophic interactions are at the heart of community ecology. Herbivores consume plants, predators consume herbivores, and pathogens and parasites infect, and sometimes kill, individuals of all species in a food web. Given the ubiquity of trophic interactions, it is no surprise that ecologists and evolutionary biologists strive to accurately characterize them. The outcome of an interaction between individuals of different species depends upon numerous factors such as the age, sex, and even phenotype of the individuals involved and the environment in which they are in. Despite this complexity, biologists often simplify an interaction down to a single number, an interaction coefficient that describes the average outcome of interactions between members of the populations of the species. Models of interacting species tend to be very simple, and interaction coefficients are often estimated from time series of population sizes of interacting species. Although biologists have long known that this approach is often approximate and sometimes unsatisfactory, work on estimating interaction strengths in more complex scenarios, and using ecological data beyond estimates of abundance, is still in its infancy. In their paper, Matthieu Paquet and Frederic Barraquand (2023) develop a demographic model of a predator and its prey. They then simulate demographic datasets that are typical of those collected by ecologists and use integrated population modelling to explore whether they can accurately retrieve the values interaction coefficients included in their model. They show that they can with good precision and accuracy. The work takes an important step in showing that accurate interaction coefficients can be estimated from the types of individual-based data that field biologists routinely collect, and it paves for future work in this area. As if often the case with exciting papers such as this, the work opens up a number of other avenues for future research. What happens as we move from demographic models of two species interacting such as those used by Paquet and Barraquand to more realistic scenarios including multiple species? How robust is the approach to incorrectly specified process or observation models, core components of integrated population modelling that require detailed knowledge of the system under study? Integrated population models have become a powerful and widely used tool in single-species population ecology. It is high time the techniques are extended to community ecology, and this work takes an important step in showing that this should and can be done. I would hope the paper is widely read and cited. References Paquet, M., & Barraquand, F. (2023). Assessing species interactions using integrated predator-prey models. EcoEvoRxiv, ver. 2 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.32942/X2RC7W | Assessing species interactions using integrated predator-prey models | Matthieu Paquet, Frederic Barraquand | <p style="text-align: justify;">Inferring the strength of species interactions from demographic data is a challenging task. The Integrated Population Modelling (IPM) approach, bringing together population counts, capture-recapture, and individual-... |  | Community ecology, Demography, Euring Conference, Food webs, Population ecology, Statistical ecology | Tim Coulson | Ilhan Özgen-Xian | 2023-01-05 17:02:22 | View |
04 Apr 2023
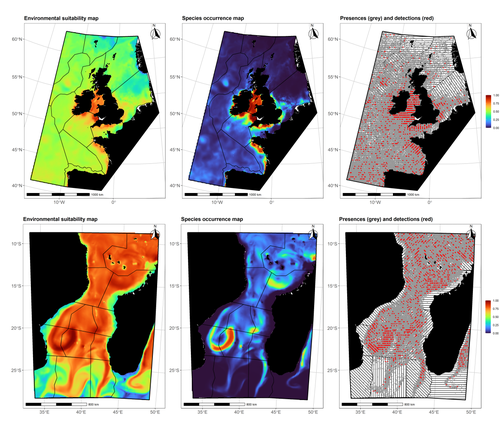
Data stochasticity and model parametrisation impact the performance of species distribution models: insights from a simulation studyCharlotte Lambert, Auriane Virgili https://doi.org/10.1101/2023.01.17.524386Species Distribution Models: the delicate balance between signal and noiseRecommended by Timothée Poisot based on reviews by Alejandra Zarzo Arias and 1 anonymous reviewer based on reviews by Alejandra Zarzo Arias and 1 anonymous reviewer
Species Distribution Models (SDMs) are one of the most commonly used tools to predict where species are, where they may be in the future, and, at times, what are the variables driving this prediction. As such, applying an SDM to a dataset is akin to making a bet: that the known occurrence data are informative, that the resolution of predictors is adequate vis-à-vis the scale at which their impact is expressed, and that the model will adequately capture the shape of the relationships between predictors and predicted occurrence. In this contribution, Lambert & Virgili (2023) perform a comprehensive assessment of different sources of complications to this process, using replicated simulations of two synthetic species. Their experimental process is interesting, in that both the data generation and the data analysis stick very close to what would happen in "real life". The use of synthetic species is particularly relevant to the assessment of SDM robustness, as they enable the design of species for which the shape of the relationship is given: in short, we know what the model should capture, and can evaluate the model performance against a ground truth that lacks uncertainty. Any simulation study is limited by the assumptions established by the investigators; when it comes to spatial data, the "shape" of the landscape, both in terms of auto-correlation and in where the predictors are available. Lambert & Virgili (2023) nicely circumvent these issues by simulating synthetic species against the empirical distribution of predictors; in other words, the species are synthetic, but the environment for which the prediction is made is real. This is an important step forward when compared to the use of e.g. neutral landscapes (With 1997), which can have statistical properties that are not representative of natural landscapes (see e.g. Halley et al., 2004). A striking point in the study by Lambert & Virgili (2023) is that they reveal a deep, indeed deeper than expected, stochasticity in SDMs; whether this is true in all models remains an open question, but does not invalidate their recommendation to the community: the interpretation of outcomes is a delicate exercise, especially because measures that inform on the goodness of the model fit do not capture the predictive quality of the model outputs. This preprint is both a call to more caution, and a call to more curiosity about the complex behavior of SDMs, while also providing a sensible template to perform future analyses of the potential issues with predictive models.
Halley, J. M., et al. (2004) “Uses and Abuses of Fractal Methodology in Ecology: Fractal Methodology in Ecology.” Ecology Letters, vol. 7, no. 3, pp. 254–71. https://doi.org/10.1111/j.1461-0248.2004.00568.x. Lambert, Charlotte, and Auriane Virgili (2023). Data Stochasticity and Model Parametrisation Impact the Performance of Species Distribution Models: Insights from a Simulation Study. bioRxiv, ver. 2 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2023.01.17.524386 With, Kimberly A. (1997) “The Application of Neutral Landscape Models in Conservation Biology. Aplicacion de Modelos de Paisaje Neutros En La Biologia de La Conservacion.” Conservation Biology, vol. 11, no. 5, pp. 1069–80. https://doi.org/10.1046/j.1523-1739.1997.96210.x. | Data stochasticity and model parametrisation impact the performance of species distribution models: insights from a simulation study | Charlotte Lambert, Auriane Virgili | <p>Species distribution models (SDM) are widely used to describe and explain how species relate to their environment, and predict their spatial distributions. As such, they are the cornerstone of most of spatial planning efforts worldwide. SDM can... |  | Biogeography, Habitat selection, Macroecology, Marine ecology, Spatial ecology, Metacommunities & Metapopulations, Species distributions, Statistical ecology | Timothée Poisot | 2023-01-20 09:43:51 | View | |
27 Nov 2023
Modeling Tick Populations: An Ecological Test Case for Gradient Boosted TreesWilliam Manley, Tam Tran, Melissa Prusinski, Dustin Brisson https://doi.org/10.1101/2023.03.13.532443Gradient Boosted Trees can deliver more than accurate ecological predictionsRecommended by Timothée Poisot based on reviews by 2 anonymous reviewers based on reviews by 2 anonymous reviewers
Tick-borne diseases are an important burden on public health all over the globe, making accurate forecasts of tick population a key ingredient in a successful public health strategy. Over long time scales, tick populations can undergo complex dynamics, as they are sensitive to many non-linear effects due to the complex relationships between ticks and the relevant (numerical) features of their environment. But luckily, capturing complex non-linear responses is a task that machine learning thrives on. In this contribution, Manley et al. (2023) explore the use of Gradient Boosted Trees to predict the distribution (presence/absence) and abundance of ticks across New York state. This is an interesting modelling challenge in and of itself, as it looks at the same ecological question as an instance of a classification problem (presence/absence) or of a regression problem (abundance). In using the same family of algorithm for both, Manley et al. (2023) provide an interesting showcase of the versatility of these techniques. But their article goes one step further, by setting up a multi-class categorical model that estimates jointly the presence and abundance of a population. I found this part of the article particularly elegant, as it provides an intermediate modelling strategy, in between having two disconnected models for distribution and abundance, and having nested models where abundance is only predicted for the present class (see e.g. Boulangeat et al., 2012, for a great description of the later). One thing that Manley et al. (2023) should be commended for is their focus on opening up the black box of machine learning techniques. I have never believed that ML models are more inherently opaque than other families of models, but the focus in this article on explainable machine learning shows how these models might, in fact, bring us closer to a phenomenological understanding of the mechanisms underpinning our observations. There is also an interesting discussion in this article, on the rate of false negatives in the different models that are being benchmarked. Although model selection often comes down to optimizing the overall quality of the confusion matrix (for distribution models, anyway), depending on the type of information we seek to extract from the model, not all types of errors are created equal. If the purpose of the model is to guide actions to control vectors of human pathogens, a false negative (predicting that the vector is absent at a site where it is actually present) is a potentially more damaging outcome, as it can lead to the vector population (and therefore, potentially, transmission) increasing unchecked. References
Boulangeat I, Gravel D, Thuiller W. Accounting for dispersal and biotic interactions to disentangle the drivers of species distributions and their abundances: The role of dispersal and biotic interactions in explaining species distributions and abundances. Ecol Lett. 2012;15: 584-593. Manley W, Tran T, Prusinski M, Brisson D. (2023) Modeling tick populations: An ecological test case for gradient boosted trees. bioRxiv, 2023.03.13.532443, ver. 3 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2023.03.13.532443 | Modeling Tick Populations: An Ecological Test Case for Gradient Boosted Trees | William Manley, Tam Tran, Melissa Prusinski, Dustin Brisson | <p style="text-align: justify;">General linear models have been the foundational statistical framework used to discover the ecological processes that explain the distribution and abundance of natural populations. Analyses of the rapidly expanding ... | Parasitology, Species distributions, Statistical ecology | Timothée Poisot | Anonymous, Anonymous | 2023-03-23 23:41:17 | View | |
28 Feb 2023
Acoustic cues and season affect mobbing responses in a bird communityAmbre Salis, Jean Paul Lena, Thierry Lengagne https://doi.org/10.1101/2022.05.05.490715Two common European songbirds elicit different community responses with their mobbing callsRecommended by Tim Parker based on reviews by 2 anonymous reviewers based on reviews by 2 anonymous reviewers
Many bird species participate in mobbing in which individuals approach a predator while producing conspicuous vocalizations (Magrath et al. 2014). Mobbing is interesting to behavioral ecologists because of the complex array of costs of benefits. Costs range from the obvious risk of approaching a predator while drawing that predator’s attention to the more mundane opportunity costs of taking time away from other activities, such as foraging. Benefits may involve driving the predator to leave, teaching relatives to recognize predators, signaling quality to conspecifics, or others. An added layer of complexity in this system comes from the inter-specific interactions that often occur among different mobbing species (Magrath et al. 2014). This study by Salis et al. (2023) explored the responses of a local bird community to mobbing calls produced by individuals of two common mobbing species in European forests, coal tits, and crested tits. Not only did they compare responses to these two different species, they assessed the impact of the number of mobbing individuals on the stimulus recordings, and they did so at two very different times of the year with different social contexts for the birds involved, winter (non-breeding) and spring (breeding). The experiment was well-designed and highly powered, and the authors tested and confirmed an important assumption of their design, and thus the results are convincing. It is clear that members of the local bird community responded differently to the two different species, and this result raises interesting questions about why these species differed in their tendency to attract additional mobbers. For instance, are species that recruit more co-mobbers more effective at recruiting because they are more reliable in their mobbing behavior (Magrath et al. 2014), more likely to reciprocate (Krams and Krama, 2002), or for some other reason? Hopefully this system, now of proven utility thanks to the current study, will be useful for following up on hypotheses such as these. Other convincing results, such as the higher rate of mobbing response in winter than in spring, also merit following up with further work. Finally, their observation that playback of vocalizations of multiple individuals often elicited a more mobbing response that the playback of vocalizations of a single individual are interesting and consistent with other recent work indicating that groups of mobbers recruit more additional mobbers than do single mobbers (Dutour et al. 2021). However, as acknowledged in the manuscript, the design of the current study did not allow a distinction between the effect of multiple individuals signaling versus an effect of a stronger stimulus. Thus, this last result leaves the question of the effect of mobbing group size in these species open to further study. REFERENCES Dutour M, Kalb N, Salis A, Randler C (2021) Number of callers may affect the response to conspecific mobbing calls in great tits (Parus major). Behavioral Ecology and Sociobiology, 75, 29. https://doi.org/10.1007/s00265-021-02969-7 Krams I, Krama T (2002) Interspecific reciprocity explains mobbing behaviour of the breeding chaffinches, Fringilla coelebs. Proceedings of the Royal Society of London. Series B: Biological Sciences, 269, 2345–2350. https://doi.org/10.1098/rspb.2002.2155 Magrath RD, Haff TM, Fallow PM, Radford AN (2015) Eavesdropping on heterospecific alarm calls: from mechanisms to consequences. Biological Reviews, 90, 560–586. https://doi.org/10.1111/brv.12122 Salis A, Lena JP, Lengagne T (2023) Acoustic cues and season affect mobbing responses in a bird community. bioRxiv, 2022.05.05.490715, ver. 5 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2022.05.05.490715 | Acoustic cues and season affect mobbing responses in a bird community | Ambre Salis, Jean Paul Lena, Thierry Lengagne | <p>Heterospecific communication is common for birds when mobbing a predator. However, joining the mob should depend on the number of callers already enrolled, as larger mobs imply lower individual risks for the newcomer. In addition, some ‘communi... | Behaviour & Ethology, Community ecology, Social structure | Tim Parker | 2022-05-06 09:29:30 | View |
MANAGING BOARD
Julia Astegiano
Tim Coulson
Anna Eklof
Dominique Gravel
François Massol
Ben Phillips
Cyrille Violle










