Direct submissions to PCI Ecology from bioRxiv.org are possible using the B2J service
Latest recommendations

| Id | Title * | Authors * ▲ | Abstract * | Picture * | Thematic fields * | Recommender | Reviewers | Submission date | |
|---|---|---|---|---|---|---|---|---|---|
24 May 2024
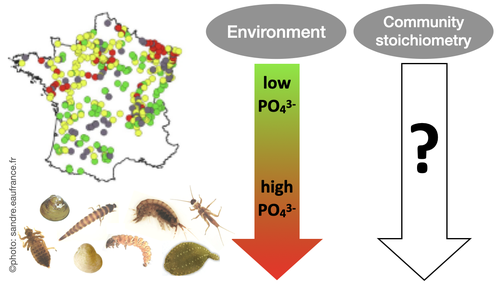
Effects of water nutrient concentrations on stream macroinvertebrate community stoichiometry: a large-scale studyMiriam Beck, Elise Billoir, Philippe Usseglio-Polatera, Albin Meyer, Edwige Gautreau, Michael Danger https://doi.org/10.1101/2024.02.01.574823The influence of water phosphorus and nitrogen loads on stream macroinvertebrate community stoichiometryRecommended by Huihuang Chen based on reviews by Thomas Guillemaud, Jun Zuo and 1 anonymous reviewer based on reviews by Thomas Guillemaud, Jun Zuo and 1 anonymous reviewer
The manuscript by Beck et al. (2024) investigates the effects of water phosphorus and nitrogen loads on stream macroinvertebrate community stoichiometry across France. Utilizing data from over 1300 standardized sampling events, this research finds that community stoichiometry is significantly influenced by water phosphorus concentration, with the strongest effects at low nitrogen levels. The results demonstrate that the assumptions of Ecological Stoichiometry Theory apply at the community level for at least two dominant taxa and across a broad spatial scale, with probable implications for nutrient cycling and ecosystem functionality. This manuscript contributes to ecological theory, particularly by extending Ecological Stoichiometry Theory to include community-level interactions, clarifying the impact of nutrient concentrations on community structure and function, and informing nutrient management and conservation strategies. In summary, this study not only addresses a gap in community-level stoichiometric research but also delivers crucial empirical support for advancing ecological science and promoting environmental stewardship. References Beck M, Billoir E, Usseglio-Polatera P, Meyer A, Gautreau E and Danger M (2024) Effects of water nutrient concentrations on stream macroinvertebrate community stoichiometry: a large-scale study. bioRxiv, 2024.02.01.574823, ver. 2 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2024.02.01.574823 | Effects of water nutrient concentrations on stream macroinvertebrate community stoichiometry: a large-scale study | Miriam Beck, Elise Billoir, Philippe Usseglio-Polatera, Albin Meyer, Edwige Gautreau, Michael Danger | <p>Basal resources generally mirror environmental nutrient concentrations in the elemental composition of their tissue, meaning that nutrient alterations can directly reach consumer level. An increased nutrient content (e.g. phosphorus) in primary... |  | Community ecology, Ecological stoichiometry | Huihuang Chen | Thomas Guillemaud, Jun Zuo, Anonymous | 2024-02-02 10:14:01 | View |
05 Feb 2020
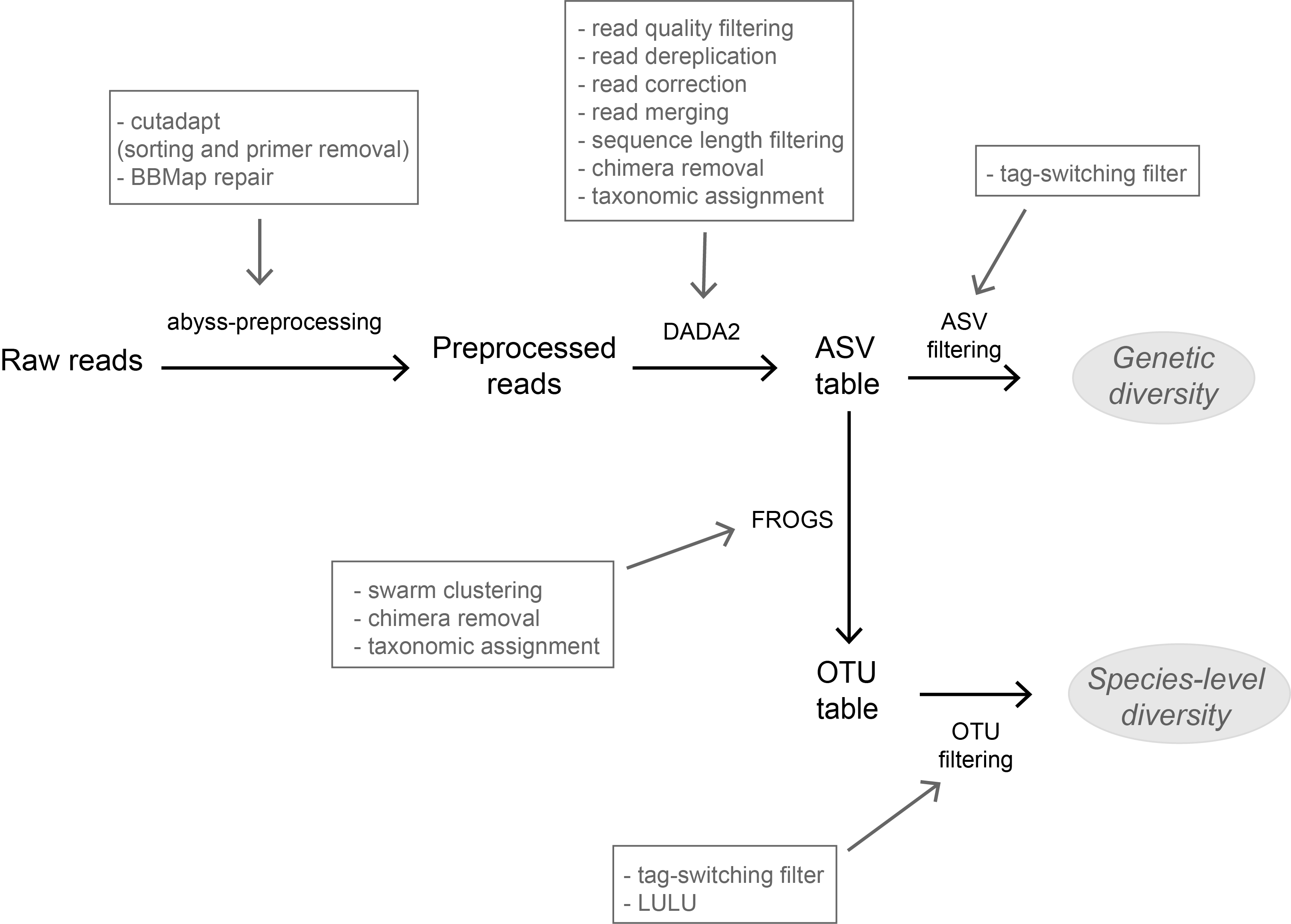
A flexible pipeline combining clustering and correction tools for prokaryotic and eukaryotic metabarcodingMiriam I Brandt, Blandine Trouche, Laure Quintric, Patrick Wincker, Julie Poulain, Sophie Arnaud-Haond https://doi.org/10.1101/717355A flexible pipeline combining clustering and correction tools for prokaryotic and eukaryotic metabarcodingRecommended by Stefaniya Kamenova based on reviews by Tiago Pereira and 1 anonymous reviewer based on reviews by Tiago Pereira and 1 anonymous reviewer
High-throughput sequencing-based techniques such as DNA metabarcoding are increasingly advocated as providing numerous benefits over morphology‐based identifications for biodiversity inventories and ecosystem biomonitoring [1]. These benefits are particularly apparent for highly-diversified and/or hardly accessible aquatic and marine environments, where simple water or sediment samples could already produce acceptably accurate biodiversity estimates based on the environmental DNA present in the samples [2,3]. However, sequence-based characterization of biodiversity comes with its own challenges. A major one resides in the capacity to disentangle true biological diversity (be it taxonomic or genetic) from artefactual diversity generated by sequence-errors accumulation during PCR and sequencing processes, or from the amplification of non-target genes (i.e. pseudo-genes). On one hand, the stringent elimination of sequence variants might lead to biodiversity underestimation through the removal of true species, or the clustering of closely-related ones. On the other hand, a more permissive sequence filtering bears the risks of biodiversity inflation. Recent studies have outlined an excellent methodological framework for addressing this issue by proposing bioinformatic tools that allow the amplicon-specific error-correction as alternative or as complement to the more arbitrary approach of clustering into Molecular Taxonomic Units (MOTUs) based on sequence dissimilarity [4,5]. But to date, the relevance of amplicon-specific error-correction tools has been demonstrated only for a limited set of taxonomic groups and gene markers. References [1] Porter, T. M., and Hajibabaei, M. (2018). Scaling up: A guide to high-throughput genomic approaches for biodiversity analysis. Molecular Ecology, 27(2), 313–338. doi: 10.1111/mec.14478 | A flexible pipeline combining clustering and correction tools for prokaryotic and eukaryotic metabarcoding | Miriam I Brandt, Blandine Trouche, Laure Quintric, Patrick Wincker, Julie Poulain, Sophie Arnaud-Haond | <p>Environmental metabarcoding is an increasingly popular tool for studying biodiversity in marine and terrestrial biomes. With sequencing costs decreasing, multiple-marker metabarcoding, spanning several branches of the tree of life, is becoming ... |  | Biodiversity, Community ecology, Marine ecology, Molecular ecology | Stefaniya Kamenova | 2019-08-02 20:52:45 | View | |
06 Dec 2019

Does phenology explain plant-pollinator interactions at different latitudes? An assessment of its explanatory power in plant-hoverfly networks in French calcareous grasslandsNatasha de Manincor, Nina Hautekeete, Yves Piquot, Bertrand Schatz, Cédric Vanappelghem, François Massol https://doi.org/10.5281/zenodo.2543768The role of phenology for determining plant-pollinator interactions along a latitudinal gradientRecommended by Anna Eklöf based on reviews by Ignasi Bartomeus, Phillip P.A. Staniczenko and 1 anonymous reviewerIncreased knowledge of what factors are determining species interactions are of major importance for our understanding of dynamics and functionality of ecological communities [1]. Currently, when ongoing temperature modifications lead to changes in species temporal and spatial limits the subject gets increasingly topical. A species phenology determines whether it thrive or survive in its environment. However, as the phenologies of different species are not necessarily equally affected by environmental changes, temporal or spatial mismatches can occur and affect the species-species interactions in the network [2] and as such the full network structure. References [1] Pascual, M., and Dunne, J. A. (Eds.). (2006). Ecological networks: linking structure to dynamics in food webs. Oxford University Press. | Does phenology explain plant-pollinator interactions at different latitudes? An assessment of its explanatory power in plant-hoverfly networks in French calcareous grasslands | Natasha de Manincor, Nina Hautekeete, Yves Piquot, Bertrand Schatz, Cédric Vanappelghem, François Massol | <p>For plant-pollinator interactions to occur, the flowering of plants and the flying period of pollinators (i.e. their phenologies) have to overlap. Yet, few models make use of this principle to predict interactions and fewer still are able to co... |  | Interaction networks, Pollination, Statistical ecology | Anna Eklöf | 2019-01-18 19:02:13 | View | |
20 Feb 2019

Differential immune gene expression associated with contemporary range expansion of two invasive rodents in SenegalNathalie Charbonnel, Maxime Galan, Caroline Tatard, Anne Loiseau, Christophe Diagne, Ambroise Dalecky, Hugues Parrinello, Stephanie Rialle, Dany Severac and Carine Brouat https://doi.org/10.1101/442160Are all the roads leading to Rome?Recommended by Simon Blanchet based on reviews by Nadia Aubin-Horth and 1 anonymous reviewerIdentifying the factors which favour the establishment and spread of non-native species in novel environments is one of the keys to predict - and hence prevent or control - biological invasions. This includes biological factors (i.e. factors associated with the invasive species themselves), and one of the prevailing hypotheses is that some species traits may explain their impressive success to establish and spread in novel environments [1]. In animals, most research studies have focused on traits associated with fecundity, age at maturity, level of affiliation to humans or dispersal ability for instance. The “composite picture” of the perfect (i.e. successful) invader that has gradually emerged is a small-bodied animal strongly affiliated to human activities with high fecundity, high dispersal ability and a super high level of plasticity. Of course, the story is not that simple, and actually a perfect invader sometimes – if not often- takes another form… Carrying on to identify what makes a species a successful invader or not is hence still an important research axis with major implications. References [1] Jeschke, J. M., & Strayer, D. L. (2006). Determinants of vertebrate invasion success in Europe and North America. Global Change Biology, 12(9), 1608-1619. doi: 10.1111/j.1365-2486.2006.01213.x | Differential immune gene expression associated with contemporary range expansion of two invasive rodents in Senegal | Nathalie Charbonnel, Maxime Galan, Caroline Tatard, Anne Loiseau, Christophe Diagne, Ambroise Dalecky, Hugues Parrinello, Stephanie Rialle, Dany Severac and Carine Brouat | <p>Background: Biological invasions are major anthropogenic changes associated with threats to biodiversity and health. What determines the successful establishment of introduced populations still remains unsolved. Here we explore the appealing as... |  | Biological invasions, Eco-immunology & Immunity, Population ecology | Simon Blanchet | 2018-10-14 12:21:52 | View | |
25 Oct 2021

The taxonomic and functional biogeographies of phytoplankton and zooplankton communities across boreal lakesNicolas F St-Gelais, Richard J Vogt, Paul A del Giorgio, Beatrix E Beisner https://doi.org/10.1101/373332The difficult interpretation of species co-distributionRecommended by Dominique Gravel based on reviews by Anthony Maire and Emilie MackeEcology is the study of the distribution of organisms in space and time and their interactions. As such, there is a tradition of studies relating abiotic environmental conditions to species distribution, while another one is concerned by the effects of consumers on the abundance of their resources. Interestingly, joining the dots appears more difficult than it would suggest: eluding the effect of species interactions on distribution remains one of the greatest challenges to elucidate nowadays (Kissling et al. 2012). Theory suggests that yes, species interactions such as predation and competition should influence range limits (Godsoe et al. 2017), but the common intuition among many biogeographers remains that over large areas such as regions and continents, environmental drivers like temperature and precipitation overwhelm their local effects. Answering this question is of primary importance in the context where species are moving around with climate warming. Inconsistencies in food web structure may arise with asynchronized movements of consumers and their resources, leading to a major disruption in regulation and potentially ecosystem functioning. Solving this problem, however, remains very challenging because we have to rely on observational data since experiments are hard to perform at the biogeographical scale. The study of St-Gelais is an interesting step forward to solve this problem. Their main objective was to assess the strength of the association between phytoplankton and zooplankton communities at a large spatial scale, looking at the spatial covariation of both taxonomic and functional composition. To do so, they undertook a massive survey of more than 100 lakes across three regions of the boreal region of Québec. Species and functional composition were recorded, along with a set of abiotic variables. Classic community ecology at this point. The difficulty they faced was to disentangle the multiple causal relationships involved in the distribution of both trophic levels. Teasing apart bottom-up and top-down forces driving the assembly of plankton communities using observational data is not an easy task. On the one hand, both trophic levels could respond to variations in temperature, nutrient availability and dissolved organic carbon. The interpretation is fairly straightforward if the two levels respond to different factors, but the situation is much more complicated when they do respond similarly. There are potentially three possible underlying scenarios. First, the phyto and zooplankton communities may share the same environmental requirements, thereby generating a joint distribution over gradients such as temperature and nutrient availability. Second, the abiotic environment could drive the distribution of the phytoplankton community, which would then propagate up and influence the distribution of the zooplankton community. Alternatively, the abiotic environment could constrain the distribution of the zooplankton, which could then affect the one of phytoplankton. In addition to all of these factors, St-Gelais et al also consider that dispersal may limit the distribution, well aware of previous studies documenting stronger dispersal limitations for zooplankton communities. Unfortunately, there is not a single statistical approach that could be taken from the shelf and used to elucidate drivers of co-distribution. Joint species distribution was once envisioned as a major step forward in this direction (Warton et al. 2015), but there are several limits preventing the direct interpretation that co-occurrence is linked to interactions (Blanchet et al. 2020). Rather, St-Gelais used a variety of multivariate statistics to reveal the structure in their observational data. First, using a Procrustes analysis (a method testing if the spatial variation of one community is correlated to the structure of another community), they found a significant correlation between phytoplankton and zooplankton communities, indicating a taxonomic coupling between the groups. Interestingly, this observation was maintained for functional composition only when interaction-related traits were considered. At this point, these results strongly suggest that interactions are involved in the correlation, but it's hard to decipher between bottom-up and top-down perspectives. A complementary analysis performed with a constrained ordination, per trophic level, provided complementary pieces of information. First observation was that only functional variation was found to be related to the different environmental variables, not taxonomic variation. Despite that trophic levels responded to water quality variables, spatial autocorrelation was more important for zooplankton communities and the two layers appear to respond to different variables. It is impossible with those results to formulate a strong conclusion about whether grazing influence the co-distribution of phytoplankton and zooplankton communities. That's the mere nature of observational data. While there is a strong spatial association between them, there are also diverging responses to the different environmental variables considered. But the contrast between taxonomic and functional composition is nonetheless informative and it seems that beyond the idiosyncrasies of species composition, trait distribution may be more informative and general. Perhaps the most original contribution of this study is the hierarchical approach to analyze the data, combined with the simultaneous analysis of taxonomic and functional distributions. Having access to a vast catalog of multivariate statistical techniques, a careful selection of analyses helps revealing key features in the data, rejecting some hypotheses and accepting others. Hopefully, we will see more and more of such multi-trophic approaches to distribution because it is now clear that the factors driving distribution are much more complicated than anticipated in more traditional analyses of community data. Biodiversity is more than a species list, it is also all of the interactions between them, influencing their distribution and abundance (Jordano 2016). References Blanchet FG, Cazelles K, Gravel D (2020) Co-occurrence is not evidence of ecological interactions. Ecology Letters, 23, 1050–1063. https://doi.org/10.1111/ele.13525 Godsoe W, Jankowski J, Holt RD, Gravel D (2017) Integrating Biogeography with Contemporary Niche Theory. Trends in Ecology & Evolution, 32, 488–499. https://doi.org/10.1016/j.tree.2017.03.008 Jordano P (2016) Chasing Ecological Interactions. PLOS Biology, 14, e1002559. https://doi.org/10.1371/journal.pbio.1002559 Kissling WD, Dormann CF, Groeneveld J, Hickler T, Kühn I, McInerny GJ, Montoya JM, Römermann C, Schiffers K, Schurr FM, Singer A, Svenning J-C, Zimmermann NE, O’Hara RB (2012) Towards novel approaches to modelling biotic interactions in multispecies assemblages at large spatial extents. Journal of Biogeography, 39, 2163–2178. https://doi.org/10.1111/j.1365-2699.2011.02663.x St-Gelais NF, Vogt RJ, Giorgio PA del, Beisner BE (2021) The taxonomic and functional biogeographies of phytoplankton and zooplankton communities across boreal lakes. bioRxiv, 373332, ver. 4 peer-reviewed and recommended by Peer community in Ecology. https://doi.org/10.1101/373332 Warton DI, Blanchet FG, O’Hara RB, Ovaskainen O, Taskinen S, Walker SC, Hui FKC (2015) So Many Variables: Joint Modeling in Community Ecology. Trends in Ecology & Evolution, 30, 766–779. https://doi.org/10.1016/j.tree.2015.09.007 Wisz MS, Pottier J, Kissling WD, Pellissier L, Lenoir J, Damgaard CF, Dormann CF, Forchhammer MC, Grytnes J-A, Guisan A, Heikkinen RK, Høye TT, Kühn I, Luoto M, Maiorano L, Nilsson M-C, Normand S, Öckinger E, Schmidt NM, Termansen M, Timmermann A, Wardle DA, Aastrup P, Svenning J-C (2013) The role of biotic interactions in shaping distributions and realised assemblages of species: implications for species distribution modelling. Biological Reviews, 88, 15–30. https://doi.org/10.1111/j.1469-185X.2012.00235.x | The taxonomic and functional biogeographies of phytoplankton and zooplankton communities across boreal lakes | Nicolas F St-Gelais, Richard J Vogt, Paul A del Giorgio, Beatrix E Beisner | <p>Strong trophic interactions link primary producers (phytoplankton) and consumers (zooplankton) in lakes. However, the influence of such interactions on the biogeographical distribution of the taxa and functional traits of planktonic organ... |  | Biogeography, Community ecology, Species distributions | Dominique Gravel | 2018-07-24 15:01:51 | View | |
31 May 2022

Sexual coercion in a natural mandrill populationNikolaos Smit, Alice Baniel, Berta Roura-Torres, Paul Amblard-Rambert, Marie J. E. Charpentier, Elise Huchard https://doi.org/10.1101/2022.02.07.479393Rare behaviours can have strong effects: evidence for sexual coercion in mandrillsRecommended by Matthieu Paquet based on reviews by Micaela Szykman Gunther and 1 anonymous reviewer based on reviews by Micaela Szykman Gunther and 1 anonymous reviewer
Sexual coercion can be defined as the use by a male of force, or threat of force, which increases the chances that a female will mate with him at a time when she is likely to be fertile, and/or decrease the chances that she will mate with other males, at some cost to the female (Smuts & Smuts 1993). It has been evidenced in a wide range of species and may play an important role in the evolution of sexual conflict and social systems. However, identifying sexual coercion in natural systems can be particularly challenging. Notably, while male behaviour may have immediate consequences on mating success (“harassment”), the mating benefits may be delayed in time (“intimidation”), and in such cases, evidencing coercion requires detailed temporal data at the individual level. Moreover, in some species male aggressive behaviours may be subtle or rare and hence hardly observed, yet still have important effects on female mating probability and fitness. Therefore, investigating the occurrence and consequences of sexual coercion in such species is particularly relevant but studying it in a statistically robust way is likely to require a considerable amount of time spent observing individuals. In this paper, Smit et al. (2022) test three clear predictions of the sexual coercion hypothesis in a natural population of Mandrills, where severe male aggression towards females is rare: (1) male aggression is more likely on sexually receptive females than on females in other reproductive states, (2) receptive females are more likely to be injured and (3) male aggression directed towards females is positively related to subsequent probability of copulation between those dyads. They also tested an alternative hypothesis, the “aggressive male phenotype” under which the correlation between male aggression towards females and subsequent mating could be statistically explained by male overall aggressivity. In agreement with the three predictions of the sexual coercion hypothesis, (1) male aggression was on average 5 times more likely, and (2) injuries twice as likely, to be observed on sexually receptive females than on females in other reproductive states and (3) copulation between males and sexually receptive females was twice more likely to be observed when aggression by this male was observed on the female before sexual receptivity. There was no support for the aggressive male hypothesis. The reviewers and I were highly positive about this study, notably regarding the way it is written and how the predictions are carefully and clearly stated, tested, interpreted, and discussed. This study is a good illustration of a case where some behaviours may not be common or obvious yet have strong effects and likely important consequences and thus be clearly worth studying. More generally, it shows once more the importance of detailed long-term studies at the individual level for our understanding of the ecology and evolution of wild populations. It is also a good illustration of the challenges faced, when comparing the likelihood of contrasting hypotheses means we need to alter sample sizes and/or the likelihood to observe at all some behaviours. For example, observing copulation within minutes after aggression (and therefore, showing statistical support for “harassment”) is inevitably less likely than observing copulations on the longer-term (and therefore showing statistical support for “intimidation”, when of course effort is put into recording such behavioural data on the long-term). Such challenges might partly explain some apparently intriguing results. For example, why are swollen females more aggressed by males if only aggression before the swollen period seems associated with more chances of mating? Here, the authors systematically provide effect sizes (and confidence intervals) and often describe the effects in an intuitive biological way (e.g., “Swollen females were, on average, about five times more likely to become injured”). This clearly helps the reader to not merely compare statistical significances but also the biological strengths of the estimated effects and the uncertainty around them. They also clearly acknowledge limits due to sample size when testing the harassment hypothesis, yet they provide precious information on the probability of observing mating (a rare behaviour) directly after aggression (already a rare behaviour!), that is, 3 times out of 38 aggressions observed between a male and a swollen female. Once again, this highlights how important it is to be able to pursue the enormous effort put so far into closely and continuously monitoring this wild population. Finally, this study raises exciting new questions, notably regarding to what extent females exhibit “counter-strategies” in response to sexual coercion, notably whether there is still scope for female mate choice under such conditions, and what are the fitness consequences of these dynamic conflicting sexual interactions. No doubt these questions will sooner than later be addressed by the authors, and I am looking forward to reading their upcoming work. References Smit N, Baniel A, Roura-Torres B, Amblard-Rambert P, Charpentier MJE, Huchard E (2022) Sexual coercion in a natural mandrill population. bioRxiv, 2022.02.07.479393, ver. 5 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2022.02.07.479393 Smuts BB, Smuts R w. (1993) Male Aggression and Sexual Coercion of Females in Nonhuman Primates and Other Mammals: Evidence and Theoretical Implications. In: Advances in the Study of Behavior (eds Slater PJB, Rosenblatt JS, Snowdon CT, Milinski M), pp. 1–63. Academic Press. https://doi.org/10.1016/S0065-3454(08)60404-0 | Sexual coercion in a natural mandrill population | Nikolaos Smit, Alice Baniel, Berta Roura-Torres, Paul Amblard-Rambert, Marie J. E. Charpentier, Elise Huchard | <p style="text-align: justify;">Increasing evidence indicates that sexual coercion is widespread. While some coercive strategies are conspicuous, such as forced copulation or sexual harassment, less is known about the ecology and evolution of inti... |  | Behaviour & Ethology | Matthieu Paquet | 2022-02-11 09:32:49 | View | |
13 May 2023

Symbiotic nutrient cycling enables the long-term survival of Aiptasia in the absence of heterotrophic food sourcesNils Radecker, Anders Meibom https://doi.org/10.1101/2022.12.07.519152Constraining the importance of heterotrophic vs autotrophic feeding in photosymbiotic cnidariansRecommended by Ulisse Cardini based on reviews by 2 anonymous reviewers based on reviews by 2 anonymous reviewers
The symbiosis with autotrophic dinoflagellate algae has enabled heterotrophic Cnidaria to thrive in nutrient-poor tropical waters (Muscatine and Porter 1977; Stanley 2006). In particular, mixotrophy, i.e. the ability to acquire nutrients through both autotrophy and heterotrophy, confers a competitive edge in oligotrophic waters, allowing photosymbiotic Cnidaria to outcompete benthic organisms limited to a single diet (e.g., McCook 2001). However, the relative importance of autotrophy vs heterotrophy in sustaining symbiotic cnidarian’s nutrition is still the subject of intense research. In fact, figuring out the cellular mechanisms by which symbiotic Cnidaria acquire a balanced diet for their metabolism and growth is relevant to our understanding of their physiology under varying environmental conditions and in response to anthropogenic perturbations. In this study's long-term starvation experiment, Radecker & Meibom (2023) investigated the survival of the photosymbiotic sea anemone Aiptasia in the absence of heterotrophic feeding. After one year of heterotrophic starvation, Apitasia anemones remained fully viable but showed an 85 % reduction in biomass. Using 13C-bicarbonate and 15N-ammonium labeling, electron microscopy and NanoSIMS imaging, the authors could clearly show that the contribution of algal-derived nutrients to the host metabolism remained unaffected as a result of increased algal photosynthesis and more efficient carbon translocation. At the same time, the absence of heterotrophic feeding caused severe nitrogen limitation in the starved Apitasia anemones. Overall, this study provides valuable insights into nutrient exchange within the symbiosis between Cnidaria and dinoflagellate algae at the cellular level and sheds new light on the importance of heterotrophic feeding as a nitrogen acquisition strategy for holobiont growth in oligotrophic waters. REFERENCES McCook L (2001) Competition between corals and algal turfs along a gradient of terrestrial influence in the nearshore central Great Barrier Reef. Coral Reefs 19:419–425. https://doi.org/10.1007/s003380000119 Muscatine L, Porter JW (1977) Reef corals: mutualistic symbioses adapted to nutrient-poor environments. Bioscience 27:454–460. https://doi.org/10.2307/1297526 Radecker N, Meibom A (2023) Symbiotic nutrient cycling enables the long-term survival of Aiptasia in the absence of heterotrophic food sources. bioRxiv, ver. 3 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2022.12.07.519152 Stanley GD Jr (2006) Photosymbiosis and the evolution of modern coral reefs. Science 312:857–858. https://doi.org/10.1126/science.1123701 | Symbiotic nutrient cycling enables the long-term survival of Aiptasia in the absence of heterotrophic food sources | Nils Radecker, Anders Meibom | <p style="text-align: justify;">Phototrophic Cnidaria are mixotrophic organisms that can complement their heterotrophic diet with nutrients assimilated by their algal endosymbionts. Metabolic models suggest that the translocation of photosynthates... |  | Eco-evolutionary dynamics, Microbial ecology & microbiology, Symbiosis | Ulisse Cardini | 2022-12-12 10:50:55 | View | |
02 Jun 2021

Identifying drivers of spatio-temporal variation in survival in four blue tit populationsOlivier Bastianelli, Alexandre Robert, Claire Doutrelant, Christophe de Franceschi, Pablo Giovannini, Anne Charmantier https://doi.org/10.1101/2021.01.28.428563Blue tits surviving in an ever-changing worldRecommended by Dieter Lukas based on reviews by Ana Sanz-Aguilar and Vicente García-Navas based on reviews by Ana Sanz-Aguilar and Vicente García-Navas
How long individuals live has a large influence on a number of biological processes, both for the individuals themselves as well as for the populations they live in. For a given species, survival is often summarized in curves showing the probability to survive from one age to the next. However, these curves often hide a large amount of variation in survival. Variation can occur from chance, or if individuals have different genotypes or phenotypes that can influence how long they might live, or if environmental conditions are not the same across time or space. Such spatiotemporal variations in the conditions that individuals experience can lead to complex patterns of evolution (Kokko et al. 2017) but because of the difficulties to obtain the relevant data they have not been studied much in natural populations. Charmantier A, Doutrelant C, Dubuc-Messier G, Fargevieille A, Szulkin M (2016) Mediterranean blue tits as a case study of local adaptation. Evolutionary Applications, 9, 135–152. https://doi.org/10.1111/eva.12282 Dubuc-Messier G, Réale D, Perret P, Charmantier A (2017) Environmental heterogeneity and population differences in blue tits personality traits. Behavioral Ecology, 28, 448–459. https://doi.org/10.1093/beheco/arw148 Kokko H, Chaturvedi A, Croll D, Fischer MC, Guillaume F, Karrenberg S, Kerr B, Rolshausen G, Stapley J (2017) Can Evolution Supply What Ecology Demands? Trends in Ecology & Evolution, 32, 187–197. https://doi.org/10.1016/j.tree.2016.12.005 Lewontin RC, Cohen D (1969) On Population Growth in a Randomly Varying Environment. Proceedings of the National Academy of Sciences, 62, 1056–1060. https://doi.org/10.1073/pnas.62.4.1056 | Identifying drivers of spatio-temporal variation in survival in four blue tit populations | Olivier Bastianelli, Alexandre Robert, Claire Doutrelant, Christophe de Franceschi, Pablo Giovannini, Anne Charmantier | <p style="text-align: justify;">In a context of rapid climate change, the influence of large-scale and local climate on population demography is increasingly scrutinized, yet studies are usually focused on one population. Demographic parameters, i... |  | Climate change, Demography, Evolutionary ecology, Life history, Population ecology | Dieter Lukas | 2021-01-29 15:24:23 | View | |
26 Apr 2021
Experimental test for local adaptation of the rosy apple aphid (Dysaphis plantaginea) during its recent rapid colonization on its cultivated apple host (Malus domestica) in EuropeOlvera-Vazquez S.G., Alhmedi A., Miñarro M., Shykoff J. A., Marchadier E., Rousselet A., Remoué C., Gardet R., Degrave A. , Robert P. , Chen X., Porcher J., Giraud T., Vander-Mijnsbrugge K., Raffoux X., Falque M., Alins, G., Didelot F., Beliën T., Dapena E., Lemarquand A. and Cornille A. https://forgemia.inra.fr/amandine.cornille/local_adaptation_dpA planned experiment on local adaptation in a host-parasite system: is adaptation to the host linked to its recent domestication?Recommended by Eric Petit based on reviews by Sharon Zytynska, Alex Stemmelen and 1 anonymous reviewerLocal adaptation shall occur whenever selective pressures vary across space and overwhelm the effects of gene flow and local extinctions (Kawecki and Ebert 2004). Because the intimate interaction that characterizes their relationship exerts a strong selective pressure on both partners, host-parasite systems represent a classical example in which local adaptation is expected from rapidly evolving parasites adapting to more evolutionary constrained hosts (Kaltz and Shykoff 1998). Such systems indeed represent a large proportion of the study-cases in local adaptation research (Runquist et al. 2020). Biotic interactions intervene in many environment-related societal challenges, so that understanding when and how local adaptation arises is important not only for understanding evolutionary dynamics but also for more applied questions such as the control of agricultural pests, biological invasions, or pathogens (Parker and Gilbert 2004). The exact conditions under which local adaptation does occur and can be detected is however still the focus of many theoretical, methodological and empirical studies (Blanquart et al. 2013, Hargreaves et al. 2020, Hoeksema and Forde 2008, Nuismer and Gandon 2008, Richardson et al. 2014). A recent review that evaluates investigations that examined the combined influence of biotic and abiotic factors on local adaptation reaches partial conclusions about their relative importance in different contexts and underlines the many traps that one has to avoid in such studies (Runquist et al. 2020). The authors of this review emphasize that one should evaluate local adaptation using wild-collected strains or populations and over multiple generations, on environmental gradients that span natural ranges of variation for both biotic and abiotic factors, in a theory-based hypothetico-deductive framework that helps interpret the outcome of experiments. These multiple targets are not easy to reach in each local adaptation experiment given the diversity of systems in which local adaptation may occur. Improving research practices may also help better understand when and where local adaptation does occur by adding controls over p-hacking, HARKing or publication bias, which is best achieved when hypotheses, date collection and analytical procedures are known before the research begins (Chambers et al. 2014). In this regard, the route taken by Olvera-Vazquez et al. (2021) is interesting. They propose to investigate whether the rosy aphid (Dysaphis plantaginea) recently adapted to its cultivated host, the apple tree (Malus domestica), and chose to pre-register their hypotheses and planned experiments on PCI Ecology (Peer Community In 2020). Though not fulfilling all criteria mentioned by Runquist et al. (2020), they clearly state five hypotheses that all relate to the local adaptation of this agricultural pest to an economically important fruit tree, and describe in details a powerful, randomized experiment, including how data will be collected and analyzed. The experimental set-up includes comparisons between three sites located along a temperature transect that also differ in local edaphic and biotic factors, and contrasts wild and domesticated apple trees that originate from the three sites and were both planted in the local, sympatric site, and transplanted to allopatric sites. Beyond enhancing our knowledge on local adaptation, this experiment will also test the general hypothesis that the rosy aphid recently adapted to Malus sp. after its domestication, a question that population genetic analyses was not able to answer (Olvera-Vazquez et al. 2020). References Blanquart F, Kaltz O, Nuismer SL, Gandon S (2013) A practical guide to measuring local adaptation. Ecology Letters, 16, 1195–1205. https://doi.org/10.1111/ele.12150 Briscoe Runquist RD, Gorton AJ, Yoder JB, Deacon NJ, Grossman JJ, Kothari S, Lyons MP, Sheth SN, Tiffin P, Moeller DA (2019) Context Dependence of Local Adaptation to Abiotic and Biotic Environments: A Quantitative and Qualitative Synthesis. The American Naturalist, 195, 412–431. https://doi.org/10.1086/707322 Chambers CD, Feredoes E, Muthukumaraswamy SD, Etchells PJ, Chambers CD, Feredoes E, Muthukumaraswamy SD, Etchells PJ (2014) Instead of “playing the game” it is time to change the rules: Registered Reports at <em>AIMS Neuroscience</em> and beyond. AIMS Neuroscience, 1, 4–17. https://doi.org/10.3934/Neuroscience.2014.1.4 Hargreaves AL, Germain RM, Bontrager M, Persi J, Angert AL (2019) Local Adaptation to Biotic Interactions: A Meta-analysis across Latitudes. The American Naturalist, 195, 395–411. https://doi.org/10.1086/707323 Hoeksema JD, Forde SE (2008) A Meta‐Analysis of Factors Affecting Local Adaptation between Interacting Species. The American Naturalist, 171, 275–290. https://doi.org/10.1086/527496 Kaltz O, Shykoff JA (1998) Local adaptation in host–parasite systems. Heredity, 81, 361–370. https://doi.org/10.1046/j.1365-2540.1998.00435.x Kawecki TJ, Ebert D (2004) Conceptual issues in local adaptation. Ecology Letters, 7, 1225–1241. https://doi.org/10.1111/j.1461-0248.2004.00684.x Nuismer SL, Gandon S (2008) Moving beyond Common‐Garden and Transplant Designs: Insight into the Causes of Local Adaptation in Species Interactions. The American Naturalist, 171, 658–668. https://doi.org/10.1086/587077 Olvera-Vazquez SG, Remoué C, Venon A, Rousselet A, Grandcolas O, Azrine M, Momont L, Galan M, Benoit L, David G, Alhmedi A, Beliën T, Alins G, Franck P, Haddioui A, Jacobsen SK, Andreev R, Simon S, Sigsgaard L, Guibert E, Tournant L, Gazel F, Mody K, Khachtib Y, Roman A, Ursu TM, Zakharov IA, Belcram H, Harry M, Roth M, Simon JC, Oram S, Ricard JM, Agnello A, Beers EH, Engelman J, Balti I, Salhi-Hannachi A, Zhang H, Tu H, Mottet C, Barrès B, Degrave A, Razmjou J, Giraud T, Falque M, Dapena E, Miñarro M, Jardillier L, Deschamps P, Jousselin E, Cornille A (2020) Large-scale geographic survey provides insights into the colonization history of a major aphid pest on its cultivated apple host in Europe, North America and North Africa. bioRxiv, 2020.12.11.421644. https://doi.org/10.1101/2020.12.11.421644 Olvera-Vazquez S.G., Alhmedi A., Miñarro M., Shykoff J. A., Marchadier E., Rousselet A., Remoué C., Gardet R., Degrave A. , Robert P. , Chen X., Porcher J., Giraud T., Vander-Mijnsbrugge K., Raffoux X., Falque M., Alins, G., Didelot F., Beliën T., Dapena E., Lemarquand A. and Cornille A. (2021) Experimental test for local adaptation of the rosy apple aphid (Dysaphis plantaginea) to its host (Malus domestica) and to its climate in Europe. In principle recommendation by Peer Community In Ecology. https://forgemia.inra.fr/amandine.cornille/local_adaptation_dp, ver. 4. Parker IM, Gilbert GS (2004) The Evolutionary Ecology of Novel Plant-Pathogen Interactions. Annual Review of Ecology, Evolution, and Systematics, 35, 675–700. https://doi.org/10.1146/annurev.ecolsys.34.011802.132339 Peer Community In. (2020, January 15). Submit your preregistration to Peer Community In for peer review. https://peercommunityin.org/2020/01/15/submit-your-preregistration-to-peer-community-in-for-peer-review/ Richardson JL, Urban MC, Bolnick DI, Skelly DK (2014) Microgeographic adaptation and the spatial scale of evolution. Trends in Ecology & Evolution, 29, 165–176. https://doi.org/10.1016/j.tree.2014.01.002 | Experimental test for local adaptation of the rosy apple aphid (Dysaphis plantaginea) during its recent rapid colonization on its cultivated apple host (Malus domestica) in Europe | Olvera-Vazquez S.G., Alhmedi A., Miñarro M., Shykoff J. A., Marchadier E., Rousselet A., Remoué C., Gardet R., Degrave A. , Robert P. , Chen X., Porcher J., Giraud T., Vander-Mijnsbrugge K., Raffoux X., Falque M., Alins, G., Didelot F., Beliën T.,... | <p style="text-align: justify;">Understanding the extent of local adaptation in natural populations and the mechanisms enabling populations to adapt to their environment is a major avenue in ecology research. Host-parasite interaction is widely se... | Evolutionary ecology, Preregistrations | Eric Petit | 2020-07-26 18:31:42 | View | ||
24 Jan 2023
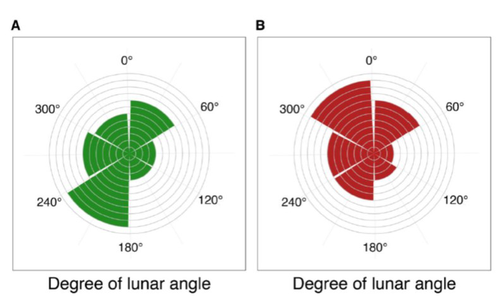
Four decades of phenology in an alpine amphibian: trends, stasis, and climatic driversOmar Lenzi, Kurt Grossenbacher, Silvia Zumbach, Beatrice Luescher, Sarah Althaus, Daniela Schmocker, Helmut Recher, Marco Thoma, Arpat Ozgul, Benedikt R. Schmidt https://doi.org/10.1101/2022.08.16.503739Alpine ecology and their dynamics under climate changeRecommended by Sergio Estay based on reviews by Nigel Yoccoz and 1 anonymous reviewerResearch about the effects of climate change on ecological communities has been abundant in the last decades. In particular, studies about the effects of climate change on mountain ecosystems have been key for understanding and communicating the consequences of this global phenomenon. Alpine regions show higher increases in warming in comparison to low-altitude ecosystems and this trend is likely to continue. This warming has caused reduced snowfall and/or changes in the duration of snow cover. For example, Notarnicola (2020) reported that 78% of the world’s mountain areas have experienced a snow cover decline since 2000. In the same vein, snow cover has decreased by 10% compared with snow coverage in the late 1960s (Walther et al., 2002) and snow cover duration has decreased at a rate of 5 days/decade (Choi et al., 2010). These changes have impacted the dynamics of high-altitude plant and animal populations. Some impacts are changes in the hibernation of animals, the length of the growing season for plants and the soil microbial composition (Chávez et al. 2021). Lenzi et al. (2023), give us an excellent study using long-term data on alpine amphibian populations. Authors show how climate change has impacted the reproductive phenology of Bufo bufo, especially the breeding season starts 30 days earlier than ~40 years ago. This earlier breeding is associated with the increasing temperatures and reduced snow cover in these alpine ecosystems. However, these changes did not occur in a linear trend but a marked acceleration was observed until mid-1990s with a later stabilization. Authors associated these nonlinear changes with complex interactions between the global trend of seasonal temperatures and site-specific conditions. Beyond the earlier breeding season, changes in phenology can have important impacts on the long-term viability of alpine populations. Complex interactions could involve positive and negative effects like harder environmental conditions for propagules, faster development of juveniles, or changes in predation pressure. This study opens new research opportunities and questions like the urgent assessment of the global impact of climate change on animal fitness. This study provides key information for the conservation of these populations. References Chávez RO, Briceño VF, Lastra JA, Harris-Pascal D, Estay SA (2021) Snow Cover and Snow Persistence Changes in the Mocho-Choshuenco Volcano (Southern Chile) Derived From 35 Years of Landsat Satellite Images. Frontiers in Ecology and Evolution, 9. https://doi.org/10.3389/fevo.2021.643850 Choi G, Robinson DA, Kang S (2010) Changing Northern Hemisphere Snow Seasons. Journal of Climate, 23, 5305–5310. https://doi.org/10.1175/2010JCLI3644.1 Lenzi O, Grossenbacher K, Zumbach S, Lüscher B, Althaus S, Schmocker D, Recher H, Thoma M, Ozgul A, Schmidt BR (2022) Four decades of phenology in an alpine amphibian: trends, stasis, and climatic drivers.bioRxiv, 2022.08.16.503739, ver. 3 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.1101/2022.08.16.503739 Notarnicola C (2020) Hotspots of snow cover changes in global mountain regions over 2000–2018. Remote Sensing of Environment, 243, 111781. https://doi.org/10.1016/j.rse.2020.111781 | Four decades of phenology in an alpine amphibian: trends, stasis, and climatic drivers | Omar Lenzi, Kurt Grossenbacher, Silvia Zumbach, Beatrice Luescher, Sarah Althaus, Daniela Schmocker, Helmut Recher, Marco Thoma, Arpat Ozgul, Benedikt R. Schmidt | <p style="text-align: justify;">Strong phenological shifts in response to changes in climatic conditions have been reported for many species, including amphibians, which are expected to breed earlier. Phenological shifts in breeding are observed i... |  | Climate change, Population ecology, Zoology | Sergio Estay | Anonymous, Nigel Yoccoz | 2022-08-18 08:25:21 | View |
MANAGING BOARD
Julia Astegiano
Tim Coulson
Anna Eklof
Dominique Gravel
François Massol
Ben Phillips
Cyrille Violle










