

 and David Alonso
and David Alonso based on reviews by 2 anonymous reviewers
based on reviews by 2 anonymous reviewers
Trophic interactions are at the heart of community ecology. Herbivores consume plants, predators consume herbivores, and pathogens and parasites infect, and sometimes kill, individuals of all species in a food web. Given the ubiquity of trophic interactions, it is no surprise that ecologists and evolutionary biologists strive to accurately characterize them.
The outcome of an interaction between individuals of different species depends upon numerous factors such as the age, sex, and even phenotype of the individuals involved and the environment in which they are in. Despite this complexity, biologists often simplify an interaction down to a single number, an interaction coefficient that describes the average outcome of interactions between members of the populations of the species. Models of interacting species tend to be very simple, and interaction coefficients are often estimated from time series of population sizes of interacting species. Although biologists have long known that this approach is often approximate and sometimes unsatisfactory, work on estimating interaction strengths in more complex scenarios, and using ecological data beyond estimates of abundance, is still in its infancy.
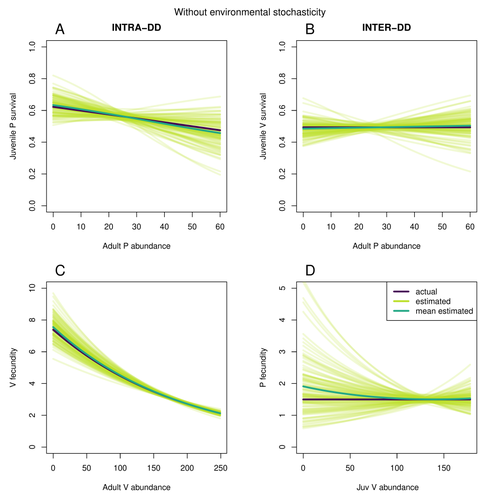
In their paper, Matthieu Paquet and Frederic Barraquand (2023) develop a demographic model of a predator and its prey. They then simulate demographic datasets that are typical of those collected by ecologists and use integrated population modelling to explore whether they can accurately retrieve the values interaction coefficients included in their model. They show that they can with good precision and accuracy. The work takes an important step in showing that accurate interaction coefficients can be estimated from the types of individual-based data that field biologists routinely collect, and it paves for future work in this area.
As if often the case with exciting papers such as this, the work opens up a number of other avenues for future research. What happens as we move from demographic models of two species interacting such as those used by Paquet and Barraquand to more realistic scenarios including multiple species? How robust is the approach to incorrectly specified process or observation models, core components of integrated population modelling that require detailed knowledge of the system under study?
Integrated population models have become a powerful and widely used tool in single-species population ecology. It is high time the techniques are extended to community ecology, and this work takes an important step in showing that this should and can be done. I would hope the paper is widely read and cited.
References
Paquet, M., & Barraquand, F. (2023). Assessing species interactions using integrated predator-prey models. EcoEvoRxiv, ver. 2 peer-reviewed and recommended by Peer Community in Ecology. https://doi.org/10.32942/X2RC7W
DOI or URL of the preprint: https://doi.org/10.32942/X2RC7W
Version of the preprint: 1
 and Tim Coulson
and Tim Coulson , posted 09 May 2023, validated 15 May 2023
, posted 09 May 2023, validated 15 May 2023We have recieved two reviews on the manuscript, and both are positive. The manuscript requires a relatively minor revision that will not require re-review -- the co-recommenders can make the call. We consequently expect to accept the revised manuscript.
The manuscript simulates data typical of those collected in a demographic field study of interacting species and then uses integrated population modelling (IPM) to recover the parameters used to simulate the data. The work is sound, and the manuscript is well written. The reviewers have identified some minor edits that will aid clarity.
One reviewer raises two slighly more major concerns. First, the reliance on a previous publication. This manuscript builds on that work, so I am not overly concerned by this. However, a few edits to make this manuscript a little more stand alone would not go amiss.
The second issue is one would hope that IPMs would recover parameters from simulations when the same model structure is used to simulate and estimate the process model in an IPM. These tests are consequently not particularly challenging. I do think this needs more acknolwedement in the discussion. In reality, we never know the generating process when we collect data, and we have to make decisions on what the process and error models will look like in an IPM. Get one of the wrong, and results can be unreliable. I am not going to request more simulations here, but in future work it might be interesting to select a broader range of models that differ from the simulation for the process model of the IPM. Perhaps stress this point more.
This simulation study investigates whether integrated population models (IPMs) with interspecific predator-prey interactions can lead to the detection of interactions, when there is actually no predator-prey interactions (or at least no demographic impact of predator-prey interactions). This work is an important milestone for the use of such IPMs to confidently evidence the demographic impact of interspecific interactions. The manuscript clearly describes the objective of the study and the chosen methodology. The results are clearly presented and convincing. I have mostly minor comments that do not challenge the validity of this study, some of them being probably due to the fact that I am not a specialist of this topic.
1-observation error
The authors use a lognormal distribution for the observation error. This generates non-integer observation values for population counts. This seems to me unrealistic. I guess that rounding to the nearest integer might probably not change much the results, or is there a reason for not doing this? (a comment on this in the method section would be sufficient to me)
2-capture-mark-recapture data
I am not sure to get the exact meaning of the m_{t,j} values presented in paragraph 2.1 (for t<j<T). Is it the number of juveniles marked at time t that have been resighted at time j for the first time (ie, not resighted before j)? In this case, maybe rewrite the equation for theta_{t,j} by putting (1-p)^{j-t}p before the product Pi, otherwise it is unclear whether this term is within this product or not.
But then, why not recording the complete history of re-sighting? For instance a juvenile marked at time t, re-sighted at time t+2 and time t+5 would appear in m_{t,t+2}, but the fact that it has been re-sighted at t+5 would not be used in the inference, while it is informative on the survival probability and on the probability of resighting. Or am I misunderstanding something, here?
3-cases without MCMC convergence
I guess that people analyzing empirical datasets and facing convergence issues would try to lengthen the chains to reach convergence. I wonder whether for such datasets the likelihood of obtaining false positives would be roughly the same or whether such problematic cases are also associated with larger probabilities of false positives?
4-model misspecification
This simulation study is performed in the favorable case in which data are simulated with the model used for the inference process (ie, no model misspecification). While this is a sensible way to proceed, I wonder whether the authors could elaborate some thoughts on the likelihood (or not) of increased occurence of false positives for certain types of model misspecification. I note that they are somehow touching on this issue in their last paragraph when recommanding to reiterate their approach to "new systems with different life history parameters and density-dependent structures". But if they could provide readers with some general (untested) predictions or warnings, this could help.
https://doi.org/10.24072/pci.ecology.100522.rev11
I think this is a very interesting paper, and it’s nice to see that IPMs might indeed be a promising method for estimating species interactions. However, I have two main concerns with the paper in its current form.
First, I wish this paper was less dependent on Barraquand & Gimenez (2019) and stood a bit more on its own. As it is, most of the methods are described on the basis of the earlier paper, focusing on how they do or don’t differ from that one and cannot be properly understood on their own. I would much rather read a thorough description of the methods used in the current paper, along with a biological justification and explanation, instead of reading that “this paper did the same as in that paper but with this small change”. This is a recurring topic, as almost every section in the methods starts with a reference to this earlier paper.
Second, and more importantly, always fitting the same IPM as was used to generate the data (except for the one case where species interactions were included in the model and not in the data) does not seem like a realistic or useful test of using IPMs to estimate species interactions. Of course, it’s a nice first step to isolate this one effect, but my worry would be that other variation in the data not being accounted for in the model would cause spurious estimates of interactions. This could be observation error (which is assumed to be known in the current paper), or environmental variation causing stochasticity in other vital rates or intraspecific density dependence, for example. Even something as simple as fitting a model without allowing for random temporal noise to the data set where such noise is included. Without such considerations I find it hard to trust that these models would actually work in a real situation.
Minor comments:
This is just a small piece of advice: the first sentence of the introduction is a bit hard to follow and rather dry. A better first sentence might make the paper more attractive to the casual reader.
L43-47: I’m not sure I follow this sentence and I don’t understand how general these results are meant to be. What is the relation to the rest of the paragraph. In short: what message are you trying to get across to the reader in this sentence?
Second paragraph in the introduction: It would be nice to link your study to studies like Mutshinda et al (2009) – Proc. Biol Sci 276 (2923-2929) where a large number of species are modeled and methods are used to identify non-zero interactions.
L122-125: It would be better to give the explanations of the notation right after the first equation in which they are used.
Equations 6-17: Please give biological interpretations of these
L168: Do you really mean i.e. (“in other words”) here?
https://doi.org/10.24072/pci.ecology.100522.rev12