

The ability to successfully sequence trace quantities of environmental DNA (eDNA) has provided unprecedented opportunities to use genetic analyses to elucidate animal ecology, behavior, and population structure without affecting the behavior, fitness, or welfare of the animal sampled. Hair associated with an animal track in the snow, the shed exoskeleton of an insect, or a swab of animal scat are all examples of non-invasive methods to collect eDNA. Despite the seemingly uncomplicated definition of "non-invasive" as proposed by Taberlet et al. [1], Lefort et al. [2] highlight that its appropriate application to sampling methods in practice is not so straightforward. For example, collecting scat left behind on the forest floor by a mammal could be invasive if feces is used by that species to mark territorial boundaries. Other collection strategies such as baited DNA traps to collect hair, capturing and handling an individual to swab or stimulate emission of a body fluid, or removal of a presumed non essential body part like a feather, fish scale, or even a leg from an insect are often described as "non-invasive" sampling methods. However, such methods cannot be considered truly non-invasive. At a minimum, attracting or capturing and handling an animal to obtain a DNA sample interrupts its normal behavioral routine, but additionally can cause both acute and long-lasting physiological and behavioral stress responses and other effects. Even invertebrates exhibit long-term hypersensitization after an injury, which manifests as heightened vigilance and enhanced escape responses [3-5].
Through an extensive analysis of 380 papers published from 2013-2018, Lefort et al. [2] document the widespread misapplication of the term "non-invasive" to methods used to sample DNA. An astonishing 58% of these papers employed the term incorrectly. A big part of the problem is that "non-invasive" is usually used by authors in the medical or veterinary sense of not breaking the skin or entering the body [6], rather than in the broader, ecological sense of Taberlet et al. [1]. The authors argue that correct use of the term matters, because it may lead naive readers – one can imagine students, policy makers, and the general public – to incorrectly assume a particular method is safe to use in a situation where disturbing the animal could affect experimental results or raise animal welfare concerns. Such assumptions can affect experimental design, as well as interpretations of one's own or others' data.
The importance of the Lefort et al. [2] paper lies in part on the authors' call for the research community to be much more careful when applying the term "non-invasive" to methods of DNA sampling. This call cannot be shrugged off as a minor problem in a few papers – as their literature review demonstrates, "non-invasive" is being applied incorrectly more often than not. The authors recognize that not all DNA sampling must be non-invasive to be useful or ethical. Examples include taking samples for DNA extraction from museum specimens, or opportunistically from carcasses of animals hunted either legally or seized by authorities from poachers. In many cases, there may be no viable non-invasive method to obtain DNA, but a researcher strives to collect samples using methods that, although they may involve taking a sample directly from the animal's body, do not disrupt, or only slightly disrupt behavior, fitness, or welfare of the animal. Thus, the other important contribution by Lefort et al. [2] is to propose the terms "non-disruptive" and "minimally-disruptive" to describe such sampling methods, which are not strictly non-invasive. While gray areas undoubtedly remain, as acknowledged by the authors, answering the call for correct use of "non-invasive" and applying the proposed new terms for certain types of invasive sampling with a focus on level of disruption, will go a long way in limiting misconceptions and misinterpretations caused by the current confusion in terminology.
References
[1] Taberlet P., Waits L. P. and Luikart G. 1999. Noninvasive genetic sampling: look before you leap. Trends Ecol. Evol. 14: 323-327. doi: 10.1016/S0169-5347(99)01637-7
[2] Lefort M.-C., Cruickshank R. H., Descovich K., Adams N. J., Barun A., Emami-Khoyi A., Ridden J., Smith V. R., Sprague R., Waterhouse B. R. and Boyer S. 2019. Blood, sweat and tears: a review of non-invasive DNA sampling. bioRxiv, 385120, ver. 4 peer-reviewed and recommended by PCI Ecology. doi: 10.1101/385120
[3] Khuong T. M., Wang Q.-P., Manion J., Oyston L. J., Lau M.-T., Towler H., Lin Y. Q. and Neely G. G. 2019. Nerve injury drives a heightened state of vigilance and neuropathic sensitization in Drosophila. Science Advances 5: eaaw4099. doi: 10.1126/sciadv.aaw4099
[4] Crook, R. J., Hanlon, R. T. and Walters, E. T. 2013. Squid have nociceptors that display widespread long-term sensitization and spontaneous activity after bodily injury. Journal of Neuroscience, 33(24), 10021-10026. doi: 10.1523/JNEUROSCI.0646-13.2013
[5] Walters E. T. 2018. Nociceptive biology of molluscs and arthropods: evolutionary clues about functions and mechanisms potentially related to pain. Frontiers in Physiololgy 9: doi: 10.3389/fphys.2018.01049
[6] Garshelis, D. L. 2006. On the allure of noninvasive genetic sampling-putting a face to the name. Ursus 17: 109-123. doi: 10.2192/1537-6176(2006)17[109:OTAONG]2.0.CO;2
DOI or URL of the preprint: https://www.biorxiv.org/content/10.1101/385120v2
Version of the preprint: 2
Dear Dr. Lefort,
Thank you for revising your pre-print, "Blood, sweat and tears...", and resubmitting to PCI Ecology for review. I returned it to the original reviewer #2, who is satisfied with most of the revisions. However, he/she still has concerns about the issue of citing papers as being out of compliance with the definition of non-invasive DNA sampling when animals were killed for methods-development purposes, and even when there was no claim of non-invasive sampling in the paper itself.
In addition, I have the following comments for you to consider when revising:
Regarding Supplementary Table 1: Sorting by first author followed by a quick visual scan, I notice there are at least 7 duplicate entries in this table: entries 48/49; 50/51; 70/71; 80/81; 83/84; 119/120; 187/188. This will change the total number of papers sampled from 342 to 335, and the percentages, proportions, etc. will have to be recalculated. Please check closely and make sure there are no additional duplicates. In addition, for the duplicate pair 80/81, the entries in the "Sampling_Method" column differ, so please revisit the paper and decide which is correct.
Also for Supp. Table 1, please include a column with the publication information for the paper (i.e., journal, volume, page numbers). And several places in the table, "Mixte" should be replaced by "Mixed."
Line 103, consider "information to draw conclusions about the specific"
Lines 124-25, consider "section of each paper"
Line 145, "Marking" should be "Mark"
Line 186, consider "When the terminology for DNA sampling is misapplied as being non-invasive when it is not,"
Line 191, consider "for judging the validity"
Line 237, should be "encounters"
Line 239, should be "effects"
Line 261, should be "on animal"
Line 286, should be "adults"
For the "Sin 7" title (line 319), please consider changing to "Equating a non-invasive procedure with non-invasive sampling". This is a more descriptive and intuitive construction than the current title.
Line 362, "reproduction" should be "reproductive"; also, should be "as proxies for"
Line 434, should be "effects"
Line 458, should be "experimentation" (i.e., not plural)
Line 467 and 475, consider "that must be" (instead of "need to" or "needs to")
Line 495, should be "animals being more attracted"
L503, consider "being indistinguishable from that of captured"
L519, should be "to obtain"
Lines 863 and 878 indicate the literature review was through May 2018, but line 865 indicates it was conducted in July 2018. Which is correct? In the method section (line 100) it says July 2018.
Line 872, consider "width of the rectangles is proportional to the number of"
Line 880, consider "width of the bars is proportional to the number of"
After making these revisions, I will examine the paper one more time and consider for formal recommendation by PCI Ecology if the revisions are satisfactory. Please let me know of any questions. I look forward to seeing your revised version.
Best Wishes, --Tom Sappington
I thank the authors for their responses to my original comments. I have only a couple of concerns that should still be addressed:
L222-228: "…did not comply with the original definition of non-invasive DNA sampling. This included … cases where animals were … even killed prior to sample collections[32]. For example, 17 of the faecal samples analysed by Kolodziej et al.[32] were obtained from the rectum of feral pigs (Sus scrofa) that had been hunted." I appreciate the authors' removing the Kierepka et al. reference in the original manuscript from this revised version. However, they have repeated the same mistake in substituting the Kolodziej et al. paper as being a case of authors claiming non-invasive DNA sampling despite killing the animal. Once again, Kolodziej et al. is a methods development paper; in this case the goal was to quantify genotyping error rates to determine reliability of future population size estimates based on non-invasive DNA samples. They approached this by comparing error rates between tissue and faecal samples taken from the same individuals, which were killed in "a driving hunt" in a forest in Germany. The paper does not indicate whether this was a hunt organized for the study, or whether the authors obtained the samples opportunistically from an independently organized hunt. Presumably the latter. Either way, the study is not relevant to the analysis here because it is about methods development, not about killing an animal to take a "non-invasive" DNA sample as implied in lines 222-228.
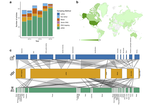
L165-166: Why is application of "non-lethal…sampling methods" to non-invasive DNA sampling "surprising"? Should not all sampling methods for non-invasive DNA sampling be "non-lethal"? Judging from Fig. 1 and Fig. 3, the category termed "non-lethal" sampling is being applied in a way that I think means taking tissue or body-part samples from a live animal without killing it. At a minimum, this term ("non-lethal") must be defined in the paper. But I would suggest finding a different term to avoid confusion; because, for example, hair from a track in the snow would be assumed by a naïve reader to be a non-lethal sampling method in the sense that collecting it does not kill the animal. If changing the term will be too disruptive because it is already a well-established term for non-lethal tissue sampling, then I don't insist, but at least define it explicitly.
And what examples are there of "lethal" sampling methods being used for "non-invasive" DNA sampling? I see a few referenced in the supplementary table 1; but given the misapplied logic of the authors in categorizing both the Kierepka et al. paper in the original (although removed in the revised version), and now the Kolodziej et al. paper in the new version as non-compliant in use of the term "non-invasive", does not instill confidence that the authors of the other papers were truly guilty of killing an animal to take a "non-invasive" sample as these categorizations and wording in the text imply. Are the authors certain these were not methods-development papers like the Kierepka et al. and Kolodziej et al. papers? I do not think the simple use of "non-invasive" in a paper that clearly describes the purpose of the study to be non-invasive methods-development should be categorized in this way just because invasive sampling (possibly including the killing of the animal) is conducted to obtain the needed samples (L271-276 revised pdf). It is perhaps ok to recommend authors be more careful in this regard to alert naïve readers who might get confused, but to classify papers like this in the literature analysis as being non-compliant with the definition of non-invasive is misleading rather than helpful.
https://doi.org/10.24072/pci.ecology.100041.rev21DOI or URL of the preprint: https://doi.org/10.1101/385120
Please see the attached response letter.
Dear Dr. Lefort,
Thank you for requesting a peer-review of your pre-print, "Blood, sweat and tears...", by PCI Ecology. I apologize for the long delay in providing the reviews, we had some difficulty finding reviewers followed by slow response from one of them.
You will see that the comments of reviewer #1 are favorable. The comments of reviewer #2 are more thorough and critical. They are essentially favorable, but suggest a major revision and restructuring of the paper before being acceptable. Although the requested revisions will require some fundamental changes in how your ideas for a different terminology in the field of DNA sampling from living animals are presented and explained, I think it would be worth the effort, and encourage you to consider doing so. If you can make such revisions, I will be happy to review the paper afresh and consider for formal recommendation by PCI Ecology.
Please let me know of any questions. I look forward to seeing your revised version. If you choose not to revise, let me know if you would, please.
Best Wishes,
--Tom Sappington
In this paper, Lefort et al. review methodology of papers claiming to use non-invasive DNA sampling over the last five years. The authors find that the methodology reported by over half of the papers reviewed do not adhere to the original definition of “non-invasive DNA sampling” proposed by Taberlet & Luikart (1999), which specifies that sampling is non-invasive when DNA is collected without disturbing or catching the animal. These results suggest that the original term has lost its meaning, and rather the term is being used to describe methods that are minimally invasive to the physical structure of the organism but might be extremely disruptive to the behavior of the organism or the organism’s fitness. Two new terms, “non-destructive sampling” and “minimally-destructive sampling” are proposed.
I found the paper very well-written, and the topic important. I believe that the methods used to conduct the review provide a representative glimpse of the recent literature. My major concern is regarding the presentation and definition of the new terms “non-destructive” and “minimally-destructive” sampling. In particular, I found Figure 2 presented contradictory information compared to what is in the text. For example, fecal sampling can be both non-invasive (sensu Taberlet) and non-disruptive if only part of the scat is collected. Or, could be considered non-disruptive when the entire scat is collected but the species is known to be non-territorial and does not use scat for social cues. Likewise, I would suspect that in many cases or for many species, remote-biopsy by darting is disruptive or at least minimally disruptive. Given the apparent gross misuse of the original term “non-invasive DNA sampling”, I think there should be no room for misinterpretation of the newly proposed terms. While Figure 2 needs work, I believe that there are also some inconsistencies in the text (e.g. Ln 402 , below), which together suggest that the new terms might need further refinement and clarification.
Ln 402: I do not agree that non-disruptive DNA sampling is required for laboratory-based experiments. Some experiments require handling or manipulation of the organism that might alter short term behavior and so taking a minimally-disruptive DNA sample at this point might have little added detrimental effect. An example of this would be when animals are collected in the wild and brought to the lab where there is a recovery and acclimation period.
https://doi.org/10.24072/pci.ecology.100041.rev11The authors are poised to make a valuable contribution to the field of eDNA sampling by suggesting better terminology to replace the use of "non-invasive" for categorizing sampling methodologies meant to not harm (or affect the behavior of) a target animal. The suggestions seem well-conceived and worthwhile. However, the way the paper is structured and conceptually focused around the results of a literature survey causes many unnecessary problems and threatens to derail the authors' main messages. I strongly suggest a thorough restructuring, as outlined at the end of my comments, where the literature survey plays only a supportive role in the authors' suggestions for terminological change. Making the survey the focus of the paper is disruptive to what should be the natural flow of the paper and the authors' arguments.
The following are comments that occurred to me as I read through the paper from beginning to end. I leave them like this, because the problems surfacing in the first half of the paper as currently written highlight the need for structural change recommended in more detail at the end.
What impact does terminology about DNA sampling potentially have on scientific behavior? The authors indicate improper use of "non-invasive" (when it actually could be disruptive) is "misleading" (Line 93). Who is being misled? The reader? The scientists themselves? What are the consequences of using "misapplied" terminology? In other words, is "misapplied" automatically the same as "misleading"? What is the hope of the authors for an outcome of using more precise terminology? Are the authors concerned that use of the term "non-invasive" DNA sampling instead of more accurate "non-disruptive" or "minimally disruptive" will open the eyes of those scientists who use such methods that they may be "invasive" after all and therefore decline to do the experiments (or to change their methodology)? It seems to me the authors need to justify the reason for conducting this study in the Introduction. If it is just to improve labeling of experimental technique for its own sake, then who cares? It would be like taking authors to task for overuse of dangling participles or something. Presumably, the authors have a more compelling reason, but they need to explain it to the reader clearly at the beginning of the paper. They approach this in lines 96-97: "This is problematic for assessing impact on animals, identifying opportunities for refinement, and for ensuring validity and quality of the data collected." But it is not clear how misapplied terminology would have these effects. One would have to assume that the scientists conducting the studies would not take their own methods (including handling of an animal for instance), whatever they may call them, into account when interpreting their data, and that this would be caused by their use of incorrect terminology. Is this what the authors mean? If so, they'll need to provide some examples, because it sounds implausible. If not, they will need to explain more precisely what they do mean.
L44: change "on the behaviour" to "on behaviour"
L71: change to "…prior to experimentation with the same individuals,…"
L80-84: So are the authors saying that the only non-invasive DNA is eDNA? That they are synonyms?
L103: Replace "(" with a comma after "noninvasive"
L107-108: the search term "ecology" is listed twice in the same string
Method section: The authors confine their description of methodology to how they conducted their literature search. However, they must also include methodology for deciding how they classified use of terminology in the papers examined. In other words, how exactly did the authors decide if a particular way of collecting DNA in a paper was non-invasive or invasive? How were categories assigned (i.e., what rules were followed), related to categories like "potentially affecting territory" or "definitely invasive" or "contact", as in Fig. 1? These rules may seem obvious to the authors, but they must be explicitly stated or described so there is no room for misunderstanding by a reader.
Section 3: "Seven deadly sins" and subheadings listing "Sin 1:", "Sin 2:", etc. ; the use of the word "sin" like this in a scientific paper seems inappropriate and will be offensive to some, probably many readers. I assume the authors mean it in a catchy, jaunty, word-play way, but I strongly recommend removing that language. It is hyperbolic with pejorative connotations, and gives the impression of prejudging the motivations of authors who have used the term "non-invasive" incorrectly up to now. Avoiding this kind of judgmental language will also improve the impact and future acceptance of the paper's ultimate recommendations for alternative, more-precise terminology by these same and other authors.
L150-153: The example of incorrect terminology is for earthworms, but the authors then state, "Yet, truly non-invasive methods exist for invertebrates" citing papers involving only arthropods (except, even more oddly, ref 19 seems to be about fish based on the title). It would seem more appropriate to use a different example of incorrect terminology based on an insect or some other arthropod rather than an annelid. Insect exuviae, pupal cases, etc. are not relevant as possible non-invasive sampling methods for earthworms, but the sentence is written as if they are.
L160-162: "It is difficult to imagine…": delete this sentence. It is subjective speculation and judgmental of motives, and is inappropriate in objective scientific writing.
L171: change "animal ware" to either "animal was" or "animals were"
L171-174: The authors' description of the Kierepka et al. (25) study is problematic and disturbing, because it misrepresents what was done in that study. They state, "For example, Kierepka et al. [25] obtained faecal samples from feral pigs (Sus scrofa) by culling the animals and squeezing faecal pellets out of the pigs’ rectum shortly after death. Such procedures clearly violate the definition proposed by Taberlet et al. [6]…" It sounded strange why anyone would be so foolish as to claim a sample was non-invasive when taken from an animal killed expressly for taking that sample, so I looked up the paper. The animal studied is an invasive pig that is controlled in DOE's Savannah River Site (SRS) to protect the habitat and native wildlife. The purpose of the study was NOT to sample the pigs non-invasively, so it is not even relevant to Taberlet's et al. definition. Instead, it was a methods development paper involving 1) development and testing of microsatellite markers that would work for this species after sampling DNA from feces; the experimental design was to compare marker amplification from actual tissue (positive control) versus from feces from the same individual of known sex, age, etc.; and 2) how well such sampling would work at time intervals after exposure of feces to the warm moist climate of the southeastern U.S. under which DNA degradation could be rapid. Kierepka et al. were developing methods for FUTURE non-invasive sampling. They never claimed that they were taking non-invasive samples in this study itself. So to characterize it that way is clearly misrepresentative, and the paper should not even be included in this study, much less used as an example of bad behavior. I cannot take the time to examine every paper the authors cite as violating "the definition of Taberlet et al." But I am concerned about how many other fundamental errors in classifying the papers there may be. Kierepka et al. is simply a paper that is irrelevant to the study undertaken in this preprint. How could the authors make such an obvious error? It is also unclear what the authors are trying to accomplish with their criticisms of misapplication of "non-invasive" terminology, as described above. I'm getting the impression it is driven at least partly by a vague sense of being critical of "hurting" other animals and misusing terminology to cover it up, even though in this case these are invasive pigs that are apparently routinely culled to protect the SRS, and even though Kierepka et al. affirm that, "All trapping and euthanasia methods conformed to the American Society of Mammalogists guidelines (Sikes et al. 2011) and University of Georgia Animal Care and Use Committee policies (Protocol 12-010-Y3-A4)." Not hurting other animals is a laudable sentiment and worthy of study. But it is all mixed up in this study to the point where the authors' immediate goal regarding their purpose is unclear, perhaps even to themselves, resulting in methodological confusion.
L207-215: How did the authors of the current paper classify these studies? Where do they fall in Fig. 1? The classification methods should be described in the Method section.
L216-219: The authors actually describe a situation like that above for the Kierepka et al. paper, where invasive sampling is used for methods development of non-invasive sampling. It is not clear why this falls under the scope of the current pre-print. Do those papers claim that the invasive sampling is non-invasive? If not, the papers do not seem relevant to the question of correct terminology.
Section 3.6: Why is this a "sin"? And who is "guilty" of it? Taberlet et al. for creating an "all or nothing" definition of non-invasive sampling? Again, the purposes of the study are muddled.
L280-281: use of the veterinary sense of non-invasive "…could minimise the perceived impacts of sampling methods on animal welfare, even if these impacts are significant in reality." Perceived by whom? The authors are either assuming the researchers in these studies using this alternative definition will not take into account animal handling when they interpret their own data, or they are worried other readers will not recognize the same, or that the researchers are covering up the possible impacts on animal welfare by using this different definition. Still not clear what the authors' concerns about impacts of use of terminology are.
The last half of the paper is much more straightforward and potentially useful to the community of researchers than the first half. My recommendation is to restructure the paper to start with a review of current methods used to take DNA samples from wildlife with the purpose, to some degree or other, of leaving the animal to carry on with its life in the wild. Then introduce the problems associated with the use of "non-invasive" terminology introduced by Taberlet et al. (including veterinary-style use of the term) as revealed by the literature analysis. Then end with the suggestions for new terminology.
In other words, the paper is most naturally a review of how DNA samples can be taken when a researcher's goal is to interfere as little as possible with the target animal's current and future well-being and activity in nature, and the pitfalls of doing this incorrectly or of making naive assumptions about impact on the animal. However, the paper currently is structured as if it is chiefly a research paper, with the analysis of terminological use of "non-invasive" in the literature as its centerpiece and most valuable aspect; and that "discovery" of misapplied terminology through this survey analysis demands a change. Focusing the paper on the literature search analysis is a mistake that makes the logic hard to follow and muddies the authors main messages which do not actually rely on the results of the analysis. These are: reminding researchers to be careful in assuming that just because they don't injure an animal when sampling doesn't mean taking the sample has no negative impact on the animal or its behavior; that even samples taken without contacting the animal (such as of fecal material) if done heedlessly could interfere with future animal behavior if that material plays an ecological or behavioral role, for example, in marking territory; and that better terminology based on more clear thinking about the effects of DNA sampling methods on the animal will help the reader more clearly and realistically evaluate a researcher's interpretation of their data. Analysis of the literature search can and should serve to buttress the authors' appeal to the scientific community to adopt better and more appropriately attributed terminology than has been used heretofore. That analysis is best presented as illustrative of the authors' main points, not the center of attention around which the rest of the paper revolves.
The appeal for better terminology as suggested by the authors seems valuable and reasonable. If the authors can restructure the paper, as well as address the many issues in the first half of the paper as described above, the authors' suggestions could have a major impact on the field. In my best judgment, leaving the paper's structure as is will probably result in little or no impact.
https://doi.org/10.24072/pci.ecology.100041.rev12