

In the recent years, morphometrics, the quantitative description of shape and its covariation [1] gained considerable momentum in evolutionary ecology. Using the form of organisms to describe, classify and try to understand their diversity can be traced back at least to Aristotle. More recently, two successive revolutions rejuvenated this idea [1–3]: first, a proper mathematical refoundation of the theory of shape, then a technical revolution in the apparatus able to acquire raw data. By using a feature extraction method and planning its massive use on data acquired by aerial drones, the study by Lacaux and colleagues [4] retraces this curse of events.
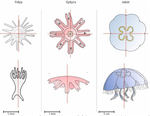
The radial symmetry of Aurelia spp. jelly fish, a common species complex, is affected by stress and more largely by environmental variations, such as pollution exposition. Aurelia spp. normally present four gonads so that the proportion of non-tetramerous individuals in a population has been proposed as a biomarker [5,6].
In this study, the authors implemented the Hough transform to largely automate the detection of the gonads in Aurelia spp. Such use of the Hough transform, a long-used approach to identify shapes through edge detection, is new to morphometrics. Here, the Aurelia spp. gonads are identified as ellipses from which aspect descriptors can be derived, and primarily counted and thus can be used to quantify the proportion of individuals presenting body plans disorders.
The sample sizes studied here were too low to allow finer-grained ecophysiological investigations. That being said, the proof-of-concept is convincing and this paper paths the way for an operational and innovative approach to the ecological monitoring of sensible aquatic ecosystems.
References
[1] Kendall, D. G. (1989). A survey of the statistical theory of shape. Statistical Science, 87-99. doi: https://doi.org/10.1214/ss/1177012589
[2] Rohlf, F. J., and Marcus, L. F. (1993). A revolution morphometrics. Trends in ecology & evolution, 8(4), 129-132. doi: https://doi.org/10.1016/0169-5347(93)90024-J
[3] Adams, D. C., Rohlf, F. J., and Slice, D. E. (2004). Geometric morphometrics: ten years of progress following the ‘revolution’. Italian Journal of Zoology, 71(1), 5-16. doi: https://doi.org/10.1080/11250000409356545
[4] Lacaux, C., Desolneux, A., Gadreaud, J., Martin-Garin, B. and Thiéry, A. (2020) Hough transform implementation to evaluate the morphological variability of the moon jellyfish (Aurelia spp.). bioRxiv, 2020.03.11.986984, ver. 3 peer-reviewed and recommended by Peer Community in Ecology. doi: https://doi.org/10.1101/2020.03.11.986984
[5] Gershwin, L. A. (1999). Clonal and population variation in jellyfish symmetry. Journal of the Marine Biological Association of the United Kingdom, 79(6), 993-1000. doi: https://doi.org/10.1017/S0025315499001228
[6] Gadreaud, J., Martin-Garin, B., Artells, E., Levard, C., Auffan, M., Barkate, A.-L. and Thiéry, A. (2017) The moon jellyfish as a new bioindicator: impact of silver nanoparticles on the morphogenesis. In: Mariottini GL, editor. Jellyfish: ecology, distribution patterns and human interactions. Nova Science Publishers; 2017. pp. 277–292.
DOI or URL of the preprint: https://www.biorxiv.org/content/10.1101/2020.03.11.986984v1
Dear Justine Gadreaud and colleagues,
The first round of revision is now over. It took some time given the lockdown and reviewers availabilities. Based on their reviews and my own, I think this preprint deserves a revision and I hope this will help.
Reviewer 1 asked for a larger dataset and for measurement error. Reviewer 2 also asked these points to be discussed. I agree with all their remarks and I also add my own comments. That being said, I think we could ultimately recommend your work, providing you answer and/or discuss these requests.
I wish you all the best,
Vincent Bonhomme
My own comments (Vincent Bonhomme)
56: I grown there, and I would have removed "often"
About the use of Matlab: perhaps you could point to open-source alternatives. eg R + magick package that binds to eponym library?
Tab2: You can perhaps remove the N= and n=
174: wonder -> tested, at least past tense everywhere? Also turns like "we continue the study" or "our interest grew" tell your story but can be made more formal.
181+185: "does not make sense" + "bilateral runs test for randomness". I'm not sure to perfectly understand and perhaps this could be made clearer with one more sentence
Overall I think that the balance could be changed a bit towards the method versus the case study you used. With such sample sizes, I'm not sure it is worth discussing so much? I have the feeling this is the main blocking point for Reviewer 1.
Also the method is well described but I think a raw output of the Hough transform on a jellyfish picture would help (eg around lines 91+)
244: "have limits" I have got the same problems myself, segmenting overlapping objects. I'm sot sure it is really limiting, I'd say it's just painful to detect and filter out such overalapping structures. Sometimes I think we could even use them but, given the number of jellyfishes during a bloom, I think the filtering is the way to go.
264: "distinguish the jellyfishes with 4 gonads and the jellyfishes with 5 gonads": if the idea is to discriminant between tetramerous and non-tetramerous why dont just use the number of detected gonads? The test you have made sounds more like a biological investigation in the how and why (as stated in lines 276+) rather than just the non-tetramerous proportion (that seems to have merits by itself) you introduced in Table 1. I think the best option would be to separate what Hough trasnforms brings here: i) first count the number of gonads, ii) allow to test biological hypotheses as you also got informative descriptors besides the number of gonads.
Additional requirements of the managing board:
As indicated in the 'How does it work?’ section and in the code of conduct, please make sure that:
-Data are available to readers, either in the text or through an open data repository such as Zenodo (free), Dryad or some other institutional repository. Data must be reusable, thus metadata or accompanying text must carefully describe the data.
-Details on quantitative analyses (e.g., data treatment and statistical scripts in R, bioinformatic pipeline scripts, etc.) and details concerning simulations (scripts, codes) are available to readers in the text, as appendices, or through an open data repository, such as Zenodo, Dryad or some other institutional repository. The scripts or codes must be carefully described so that they can be reused.
-Details on experimental procedures are available to readers in the text or as appendices.
-Authors have no financial conflict of interest relating to the article. The article must contain a "Conflict of interest disclosure" paragraph before the reference section containing this sentence: "The authors of this preprint declare that they have no financial conflict of interest with the content of this article." If appropriate, this disclosure may be completed by a sentence indicating that some of the authors are PCI recommenders: “XXX is one of the PCI XXX recommenders.”
This paper shows that the Hough transform can be applied to Jellyfish images for extracting some shape parameters relative to gonad and umbrella morphology. It is an interesting paper as the Hough transform is not (yet) a method largely implemented in biology for image detection or for getting morphometric parameters.
RM1: The method requires some input from the user (ie defining ranges for some parameter values); the author should explain how they defined these ranges (and how long it took).
The method has some potential for automatic detection, however I wonder whether it is going faster than manual digitization and application of more traditional morphometric methods such as elliptic Fourier analysis. I do not think it would be necessary to compare both approaches because they are different, but it would be good to know how long it took to extract information from the 19 images to see whether this kind of procedure could be used for very large image dataset.
RM2: In the Matlab algo, you should correct canny ege -> canny edge.
RM3: Since most of the readership here is from the ecology background, it would be good to remind in more details the hough transform method.
RM4: There is a problem of pseudoreplication and misleading information for the relative distance of the gonad to the center. As you have done after, you should average your data for each jelly fish, indeed you test for differences among jellyfishes, not among gonads. I do not see clearly why you introduce the image related to jelly fish bloom, it does not seem to be directly related with the main objective of the paper.
RM5: Finally, and I think this should be discussed, we have no idea about variation within an individual and the variation introduced by the operator (ie way to take the photography...): what if the orientation of the jelly fish is different under the camera, I guess that ellipses parameters will vary ? As the umbrella contract and dilate during the life of the animal, what about the variability of the computed parameters ? I think giving an idea of percentage of error measurement is important here because you introduce a new method, we need to now whether there is some precision and repeatability (see paper of Yezerinac et al., 1992, Systematic biology for methods regarding the estimation of error).
This being said, I definitely think that this paper is interesting because it popularize an old methodology not enough well known in ecology. I would recommend to the author to pay attention to individual variation as it might challenge the test performed here (cf the small sample size).
This is a signed recommendation: Julien CLAUDE, Institut des Sciences de l'Evolution de Montpellier. Université de Montpellier
https://doi.org/10.24072/pci.ecology.100090.rev11In this study the authors present a new image analysis and morphometric method allowing to simply and automatically assess some morphological parameters of the jellyfish Aurelia sp. They apply their method to a small sample of Jelly fish, contrasting tetramerous and non tetramerous individuals. They detect a difference in gonads position between groups as well as a difference in variability. They suggest this new tool might open up new avenues of research, in particular in ecotoxicology.
Overall, I found that the paper was clear and quite well written. The image analysis and morphometric treatments seem correctly performed and appropriate.
However, I think a larger sample size is definitely required if the authors want to tell anything on the difference among the symmetry groups and its ecological meaning. Therefore, the scope of the study – and thus of the paper – is very narrow, limited to the image analysis part. The ecological interpretation is extremely tentative.
An analysis of measurement error is also lacking, while clearly important for this kind of study.
I would thus suggest rejection.
I also have a few comments following the text.
I found it surprising that the studied species is not identified..? This should be explained.
The sample size is extremely low to infer any morphometric difference among groups.
No detail on the statistical tests applied is provided in the material and methods section. Such details are surprisingly given in the results section… The two sections, material and method and results sections are mixed up.
The rationale for the bilateral runs tests applied should be given.
It is a bit weird that the authors give the formula of the variance L226, as if it was an original statistics… just referring to the variance is fine.
An analysis of measurement error, including the effect of jelly fish positionning and picture acquisition should be performed. This is particularly important as the aim of the present study is to apply this morphometric analysis to unstandardized images – as far as I understood correctly. Even on standardized images, the ME analysis is important to assess its amount relative to the biological signal.
This is even more important as the authors are interested in relative variability of the gonads within individuals. For example, testing any paralax effect on the estimation of such variability is needed.
https://doi.org/10.24072/pci.ecology.100090.rev12